May. 09, 2011
Search and rescue missions have followed each of the devastating earthquakes that hit Haiti, New Zealand and Japan during the past 18 months. Machines able to navigate through complex dirt and rubble environments could have helped rescuers after these natural disasters, but building such machines is challenging.
Researchers at the Georgia Institute of Technology recently built a robot that can penetrate and "swim" through granular material. In a new study, they show that varying the shape or adjusting the inclination of the robot's head affects the robot's movement in complex environments.
"We discovered that by changing the shape of the sand-swimming robot's head or by tilting its head up and down slightly, we could control the robot's vertical motion as it swam forward within a granular medium,” said Daniel Goldman, an assistant professor in the Georgia Tech School of Physics.
Results of the study will be presented on May 10 at the 2011 IEEE International Conference on Robotics and Automation in Shanghai. Funding for this research was provided by the Burroughs Wellcome Fund, National Science Foundation and Army Research Laboratory.
The study was conducted by Goldman, bioengineering doctoral graduate Ryan Maladen, physics graduate student Yang Ding and physics undergraduate student Andrew Masse, all from Georgia Tech, and Northwestern University mechanical engineering adjunct professor Paul Umbanhowar.
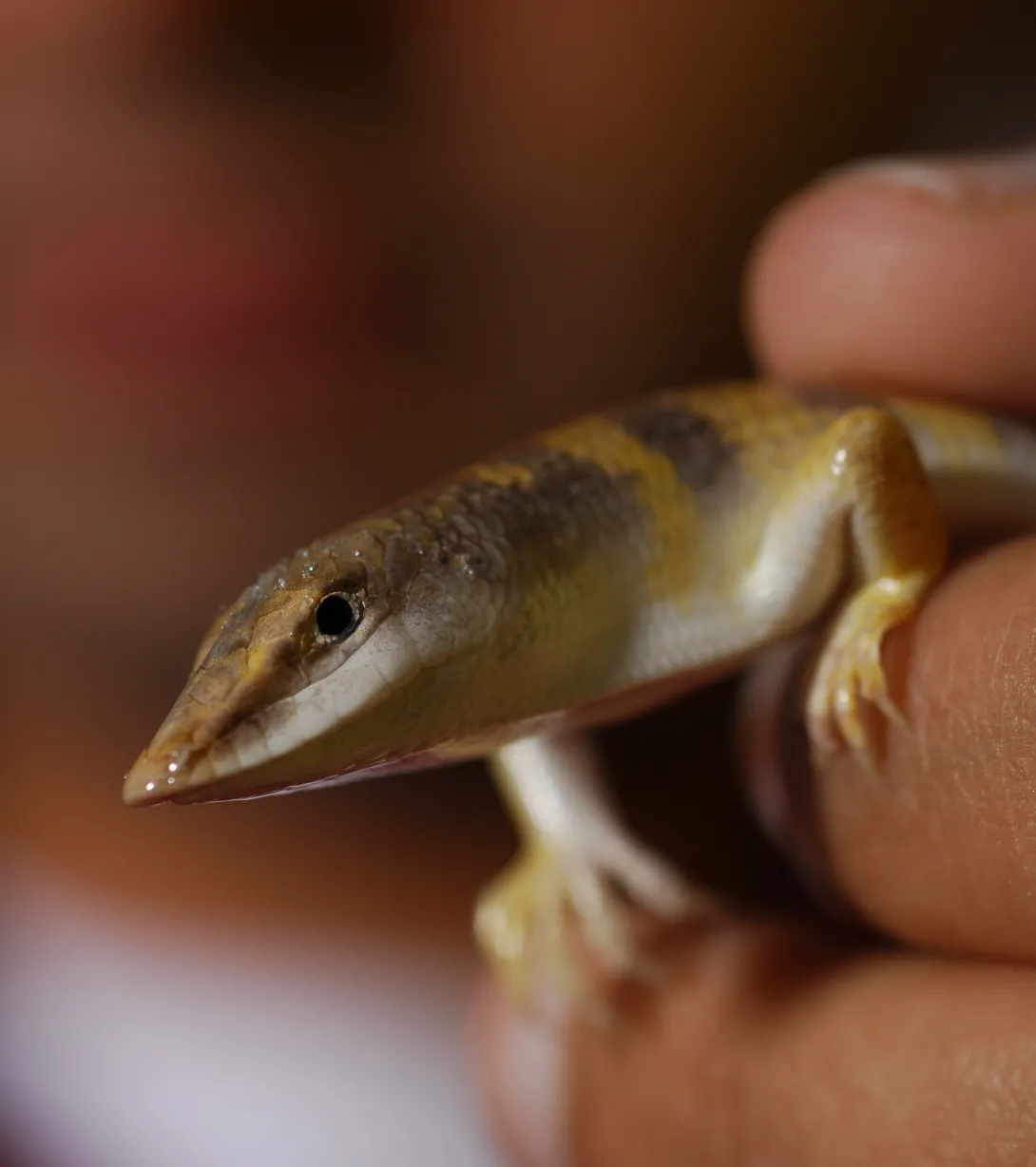
"The biological inspiration for our sand-swimming robot is the sandfish lizard, which inhabits the Sahara desert in Africa and rapidly buries into and swims within sand," explained Goldman. "We were intrigued by the sandfish lizard's wedge-shaped head that forms an angle of 140 degrees with the horizontal plane, and we thought its head might be responsible for or be contributing to the animal's ability to maneuver in complex environments."
For their experiments, the researchers attached a wedge-shaped block of wood to the head of their robot, which was built with seven connected segments, powered by servo motors, packed in a latex sock and wrapped in a spandex swimsuit. The doorstop-shaped head -- which resembled the sandfish's head -- had a fixed lower length of approximately 4 inches, height of 2 inches and a tapered snout. The researchers examined whether the robot's vertical motion could be controlled simply by varying the inclination of the robot's head.
Before each experimental run in a test chamber filled with quarter-inch-diameter plastic spheres, the researchers submerged the robot a couple inches into the granular medium and leveled the surface. Then they tracked the robot's position until it reached the end of the container or swam to the surface.
The researchers investigated the vertical movement of the robot when its head was placed at five different degrees of inclination. They found that when the sandfish-inspired head with a leading edge that formed an angle of 155 degrees with the horizontal plane was set flat, negative lift force was generated and the robot moved downward into the media. As the tip of the head was raised from zero to 7 degrees relative to the horizontal, the lift force increased until it became zero. At inclines above 7 degrees, the robot rose out of the medium.
"The ability to control the vertical position of the robot by modulating its head inclination opens up avenues for further research into developing robots more capable of maneuvering in complex environments, like debris-filled areas produced by an earthquake or landslide," noted Goldman.
The robotics results matched the research team's findings from physics experiments and computational models designed to explore how head shape affects lift in granular media.
"While the lift forces of objects in air, such as airplanes, are well understood, our investigations into the lift forces of objects in granular media are some of the first ever," added Goldman.
For the physics experiments, the researchers dragged wedge-shaped blocks through a granular medium. Blocks with leading edges that formed angles with the horizontal plane of less than 90 degrees resembled upside-down doorstops, the block with a leading edge equal to 90 degrees was a square, and blocks with leading edges greater than 90 degrees resembled regular doorstops.
They found that blocks with leading edges that formed angles with the horizontal plane less than 80 degrees generated positive lift forces and wedges with leading edges greater than 120 degrees created negative lift. With leading edges between 80 and 120 degrees, the wedges did not generate vertical forces in the positive or negative direction.
Using a numerical simulation of object drag and building on the group’s previous studies of lift and drag on flat plates in granular media, the researchers were able to describe the mechanism of force generation in detail.
"When the leading edge of the robot head was less than 90 degrees, the robot's head experienced a lift force as it moved forward, which resulted in a torque imbalance that caused the robot to pitch and rise to the surface," explained Goldman.
Since this study, the researchers have attached a wedge-shaped head on the robot that can be dynamically modulated to specific angles. With this improvement, the researchers found that the direction of movement of the robot is sensitive to slight changes in orientation of the head, further validating the results from their physics experiments and computational models.
Being able to precisely control the tilt of the head will allow the researchers to implement different strategies of head movement during burial and determine the best way to wiggle deep into sand. The researchers also plan to test the robot's ability to maneuver through material similar to the debris found after natural disasters and plan to examine whether the sandfish lizard adjusts its head inclination to ensure a straight motion as it dives into the sand.
This material is based on research sponsored by the Burroughs Wellcome Fund, the National Science Foundation (NSF) under Award Number PHY-0749991, and the Army Research Laboratory (ARL) under Cooperative Agreement Number W911NF-08-2-0004. The views and conclusions contained in this document are those of the authors and should not be interpreted as representing the official policies, either expressed or implied, of NSF, ARL or the U.S. government.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364
May. 02, 2011
The National Institutes of Health (NIH) has awarded nearly $5.4 million to a collaborative team of scientists at Emory University, Georgia Institute of Technology and Texas Biomedical Research Institute to develop vaccines for prevention of deadly hemorrhagic fevers.
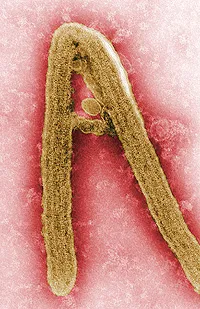
The project focuses on the development of an effective vaccine for Ebola and Marburg virus infections, two members of a family named "filoviruses" because they produce long filamentous particles.
The lead investigators include Richard Compans and Chinglai Yang at Emory University, Mark Prausnitz at Georgia Tech, and Jean Patterson and Ricardo Carrion at Texas Biomedical Research Institute.
According to Compans, "These viruses cause severe hemorrhagic fevers with up to 90 percent mortality, and can be passed via person-to-person contact, thus posing a high risk in case of an epidemic outbreak as well as a possible bioterrorism threat.”
In ongoing research, the Emory group has developed virus-like particle (VLP) vaccines to prevent virus infection, and has shown that the Ebola VLPs stimulate immune cell activity and induce strong antibody responses, indicating that such VLPs could be effective vaccines to induce protective immunity against infection. They also have found that immunization with a mixture of DNA and VLP vaccines (DNA/VLP) induced higher levels of protective immune responses in comparison to immunization with either vaccine alone.
"We consider this to be one of the most promising and safest approaches to protecting against hemorrhagic fever viruses," said Patterson, chair of the Department of Virology and Immunology at Texas Biomedical Research Institute.
In addition, the researchers are testing these vaccines with a new skin delivery technology developed at Georgia Tech that could further increase such responses, with the aim of having a vaccine that can confer rapid and long-lasting protection against Ebola and Marburg virus infection. The results will identify the most effective candidate vaccine for human trials. The successful development of this vaccine strategy may also lead to vaccines against other viral hemorrhagic fevers, which still lack effective vaccines.
"Administering these vaccines with a microneedle skin patch may increase the effectiveness of the vaccine, as well as potentially make vaccination simple and painless," said Prausnitz, professor of chemical and biomedical engineering at Georgia Tech.
The Robert W. Woodruff Health Sciences Center of Emory University produced this news release.
Media Relations Contacts: Holly Korschun, Emory University (hkorsch@emory.edu)(404-727-3990) or John Toon, Georgia Tech (jtoon@gatech.edu)(404-894-6986).
News Contact
John Toon
Research News & Publications Office
Contact John Toon
404-894-6986
Apr. 04, 2011
A new study reveals that a group of ancient enzymes adapted to substantial changes in ocean temperature and acidity during the last four billion years, providing evidence that life on Early Earth evolved from a much hotter, more acidic environment to the cooler, less acidic global environment that exists today.
The study found that a group of ancient enzymes known as thioredoxin were chemically stable at temperatures up to 32 degrees Celsius (58 degrees Fahrenheit) higher than their modern counterparts. The enzymes, which were several billion years old, also showed increased activity at lower pH levels -- which correspond to greater acidity.
"This study shows that a group of ubiquitous proteins operated in a hot, acidic environment during early life, which supports the view that the environment progressively cooled and became more alkaline between four billion and 500 million years ago," said Eric Gaucher, an associate professor in the School of Biology at the Georgia Institute of Technology.
The study, which was published April 3 in the advance online edition of the journal Nature Structural & Molecular Biology, was conducted by an international team of researchers from Georgia Tech, Columbia University and the Universidad de Granada in Spain.
Major funding for this study was provided by two grants from the National Aeronautics and Space Administration to Georgia Tech, a grant from the National Institutes of Health to Columbia University, and a grant from the Spanish Ministry of Science and Innovation to the Universidad de Granada.
Using a technique called ancestral sequence reconstruction, Gaucher and Georgia Tech biology graduate student Zi-Ming Zhao reconstructed seven ancient thioredoxin enzymes from the three domains of life -- archaea, bacteria and eukaryote -- that date back between one and four billion years.
To resurrect these enzymes, which are found in nearly all known modern organisms and are essential for life in mammals, the researchers first constructed a family tree of the more than 200 thioredoxin sequences available from the three domains of life. Then they reconstructed the sequences of the ancestral thioredoxin enzymes using statistical methods based on maximum likelihood. Finally, they synthesized the genes that encoded these sequences, expressed the ancient proteins in the cells of modern Escherichia coli bacteria and then purified the proteins.
"By resurrecting proteins, we are able to gather valuable information about the adaptation of extinct forms of life to climatic, ecological and physiological alterations that cannot be uncovered through fossil record examinations," said Gaucher.
The reconstructed enzymes from the Precambrian period -- which ended about 542 million years ago -- were used to examine how environmental conditions, including pH and temperature, affected the evolution of the enzymes and their chemical mechanisms.
"Given the ancient origin of the reconstructed thioredoxin enzymes, with some of them predating the buildup of atmospheric oxygen, we thought their catalytic chemistry would be simple, but we found that thioredoxin enzymes use a complex mixture of chemical mechanisms that increases their efficiency over the simpler compounds that were available in early geochemistry," said Julio Fernández, a professor in the Department of Biological Sciences professor at Columbia University.
Fernández led a team that included Columbia University postdoctoral researchers Raul Perez-Jimenez, Jorge Alegre-Cebollada and Sergi Garcia-Manyes, and graduate student Pallav Kosuri in using an assay based on single molecule force spectroscopy to measure the activity level of the thioredoxin enzymes under different pH levels.
For their experiments, the researchers used an atomic force microscope to pick up and stretch an engineered protein in a solution containing thioredoxin. They first applied a constant force to the protein, causing it to rapidly unfold and expose its disulfide bonds to the thioredoxin enzymes. The rate at which a thioredoxin enzyme snipped the disulfide bonds determined the enzyme's level of efficiency.
The study results showed that the three oldest thioredoxin enzymes -- those thought to have inhabited Earth 4.2 to 3.5 billion years ago -- were able to operate in lower pH environments than the modern thioredoxin enzymes.
"Our analysis indicates that ancient thioredoxin enzymes were well adapted to function under acidic conditions and that they maintained their high level of activity as they evolved in more alkaline environments," said Fernández.
To measure the temperature range in which the enzymes operated, professor Jose Sanchez-Ruiz and graduate student Alvaro Inglés-Prieto from the Departamento de Química-Física at the Universidad de Granada in Spain used a technique called differential scanning calorimetry. This method measures the stability of enzymes by heating the enzymes at a constant rate and measuring the heat change associated with their unfolding.
The researchers found that the ancient proteins were stable at temperatures up to 32 degrees Celsius higher than the modern thioredoxins. The experiments showed that the enzymes exhibited higher temperature stability the older they were. The results provide evidence that ancestral thioredoxins adapted to the cooling trend of ancient oceans, as inferred from geological records.
"Our results confirm that life has the remarkable ability to adapt to a wide range of historical environmental conditions; and by extension, life will undoubtedly adapt to future environmental changes, albeit at some cost to many species," said Gaucher.
This study also showed that the experimental resurrection of ancient proteins together with the sensitivity of single-molecule techniques can be a powerful tool for understanding the origin and evolution of life on Earth.
The researchers are currently using this strategy to assess other enzymes to get a clearer picture of what life was like on Early Earth. They are also applying these tools to the field of biotechnology, where enzymes play important roles in many industrial processes.
"The functions and characteristics we observed in the ancestral enzymes show that our techniques can be implemented to generate improved enzymes for a wide range of applications," added Perez-Jimenez.
This project was supported by the National Aeronautics and Space Administration (NASA) (Award Nos. NNX08AO12G and NNA09DA78A). The content is solely the responsibility of the principal investigator and does not necessarily represent the official view of NASA.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364
Mar. 28, 2011
Physicians and engineerswithin a new center devoted to pediatric nanomedicine will develop targeted,molecular-sized nanoparticles as part of a unique approach to treatingpediatric diseases. Specific focus areas will include pediatric heart diseaseand thrombosis, infectious diseases, cancer, sickle cell disease and cysticfibrosis.
The Center for PediatricNanomedicine (CPN) is the first of its kind in the world.
Directed by Gang Bao, thecenter will involve researchers from Emory University, the Georgia Institute ofTechnology and Children’s Healthcare of Atlanta.
“Because nano-scalestructures are compatible in size to biomolecules, nanomedicine providesunprecedented opportunities for achieving better control of biologicalprocesses and drastic improvements in disease detection, therapy andprevention,” says Bao, the Robert A. Milton Professor of Biomedical Engineeringin the Wallace H. Coulter Department of Biomedical Engineering at Georgia Techand Emory University.
Nanomedicine involves thedevelopment of engineered nanoscale structures and devices for betterdiagnostics and highly specific medical interventions to treat diseases andrepair damaged tissues. One nanometer is one-billionth of a meter.
The CPN is part of theEmory-Children’s Pediatric Research Center led by the two institutions,including partnerships with Georgia Institute of Technology and Morehouse Schoolof Medicine.
With the leadership of Dr. PaulSpearman, Children’s chief research officer and vice chair for research in theEmory University Department of Pediatrics, 14 key priority centers have beenidentified. These are hematology and oncology; immunology and vaccines;transplant immunology and immune therapeutics; pediatric healthcare technologyinnovation; cystic fibrosis; developmental lung biology; endothelial biology;cardiovascular biology; drug discovery; autism; neurosciences; nanomedicine; outcomesresearch and public health; and clinical and translational research.
Emory and Georgia Techalready have had significant and successful research partnerships innanomedicine funded by the National Institutes of Health. These have includednanotechnology center of excellence for the detection and treatment ofcardiovascular disease, the development of personalized and predictiveoncology, and the development of engineered protein machines for treatingsingle-gene disorders.
“Nanotechnology can beapplied to many diseases, and the application of nanotechnology could have aprofound impact on improving children’s health,” says Bao.
Current centers located inthe joint Georgia Tech-Emory biomedical engineering department include theCenter for Translational Cardiovascular Nanomedicine (funded by a $14.6million, five-year grant from NHLBI/NIH) and the Nanomedicine Center forNucleoprotein Machines (funded by a $16.1 million, five-year grant from NIH).
The discoveries made inthese centers also will be applied to research in pediatric diseases. Forexample, scientists in the center for nucleoprotein machines are focused ondeveloping a technology to correct single-gene defects that lead to humandisease. They hope to use this approach to treat and eventually cure sicklecell disease, first focusing on curing a mouse model of sickle cell. The newtechnology would then be applied to human sickle cell patients.
“Nanomedicine is expected todramatically exceed what has occurred in the field thus far, and our belief isthat it will revolutionize medicine,” says Bao. “We plan to make this newpediatric nanomedicine center a leader in applying these unique discoveries totreating and curing children’s diseases.”
The biomedical engineeringfaculty members who are involved in the CPN activities include: Dr. WilburLam, biomedical engineer; Barbara Boyan, professor and Price Gilbert Jr. Chairin Tissue Engineering and associate dean for research; Niren Murthy, associateprofessor of biomedical engineering; Michael Davis, assistant professor ofbiomedical engineering; Phil Santangelo, assistant professor of biomedicalengineering; Shuming Nie, professor and the Wallace H. Coulter DistinguishedFaculty Chair in Biomedical Engineering; Thomas Barker, assistant professor ofbiomedical engineering; and Ravi Bellamkonda, professor and associate vicepresident for research.
News Contact
Georgia Tech Media Relations
Laura Diamond
laura.diamond@comm.gatech.edu
404-894-6016
Jason Maderer
maderer@gatech.edu
404-660-2926
Mar. 23, 2011
Avian embryos could join the list of model organisms used to study a specific type of cell migration called epiboly, thanks to the results of a study published this month in the journal Developmental Dynamics. The new study provides insights into the mechanisms of epiboly, a developmental process involving mass movement of cells as a sheet, which is linked with medical conditions that include wound healing and cancer.
The study, published online on March 15, explains how epithelial cells expand as a sheet and migrate to engulf the entire avian egg yolk as it grows. It also reveals the presence of certain molecules during this process that have not been previously reported in other major developmental models, including Xenopus frogs and zebrafish.
"These molecules and mechanisms of early development in the avian embryo may demonstrate evolutionary differences across species in the collective movement of epithelial cells and motivate additional studies of avian embryo development," said Evan Zamir, an assistant professor in the George W. Woodruff School of Mechanical Engineering at Georgia Tech.
Matt Futterman, who worked on the project as a graduate student at Georgia Tech, and mechanical engineering professor Andrés García also contributed to this study. The research was funded by Zamir's new faculty support from Georgia Tech and by a grant to García from the National Institutes of Health.
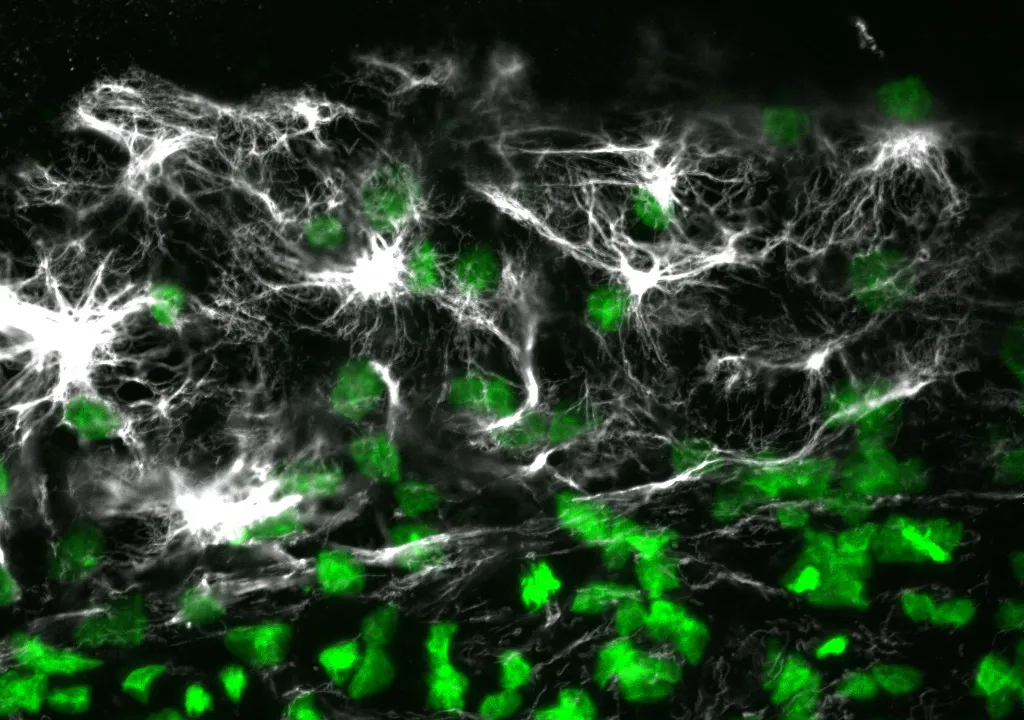
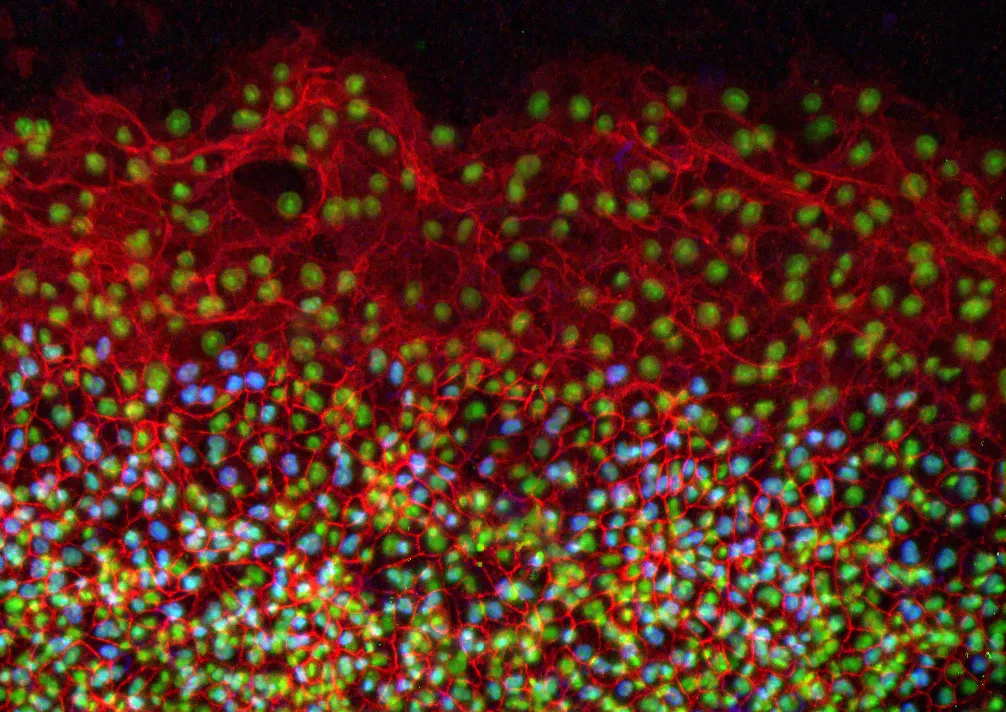
In the study, the researchers conducted immunofluorescence and high-resolution confocal microscopy experiments to examine the spatial distribution and expression of five proteins -- vimentin, cytokeratin, β-catenin, E-cadherin and laminin -- as cells moved to wrap the yolk sac of quail embryos during development.
The results showed that during this process, four of the proteins -- vimentin, cytokeratin, β-catenin and E-cadherin -- appeared in the cells located at the free edge of the migrating cell sheet. Finding dense interconnected networks of both vimentin and cytokeratin in the edge cells surprised the researchers.
"Since cytokeratin is generally associated with the epithelial phenotype and vimentin is generally associated with the mesenchymal phenotype, it's rare to see them expressed in the same cells, but this does occur in metastasizing tumor cells," said Zamir.
Cells expressing the mesenchymal phenotype are typically found in connective tissues -- such as bone, cartilage, and the lymphatic and circulatory systems -- whereas cells of the epithelial phenotype are found in cavities and glands and on surfaces throughout the body.
This finding provides evidence that epithelial cells normally attached to a membrane surface underwent biochemical changes that enabled them to assume a mesenchymal cell phenotype, which enhanced their migratory capacity. This process, called partial epithelial-to-mesenchymal transition, has many similarities to the initiation of tumor cell metastasis and wound healing.
Since this epithelial and mesenchymal expression pattern in the edge cells has not previously been reported in Xenopus or zebrafish, it may be unique to the avian embryo. This discovery would make the avian embryo a valuable model for studying tumor cell migration and wound healing.
In addition to detailing protein expression in the quail embryo during development, the researchers also determined the origin of the new cells required at the migrating edge to cover the growing yolk. During development, the radius of the quail yolk doubles every day for the first few days, representing a hundreds-fold increase in the egg yolk surface area.
"For each individual cell that has to cover the egg yolk as it grows, the migration around the yolk is extraordinary, because it's such a large territory -- it would be like an ant walking across the earth," explained Zamir.
Looking more closely at the edge cells, the researchers found strong evidence that expansion of the edge cell population was due exclusively to cells relocating from an interior region to the edge as the embryo expanded. The cells located at the free edge generated the bulk of the traction force necessary for expansion and towed the cells within the interior of the epithelium.
"These experiments confirm that edge cell proliferation is not the primary mechanism for expansion of the edge cell population," noted Zamir. "And our observation of epithelial-to-mesenchymal transition in the edge cells explains how these epithelial cells might be changing phenotype to become migratory in this rapidly expanding sheet."
To determine if this study's findings are indeed unique to the avian embryo, Zamir plans to conduct further studies to characterize protein expression and cell migration in Xenopus and zebrafish.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364
Mar. 02, 2011
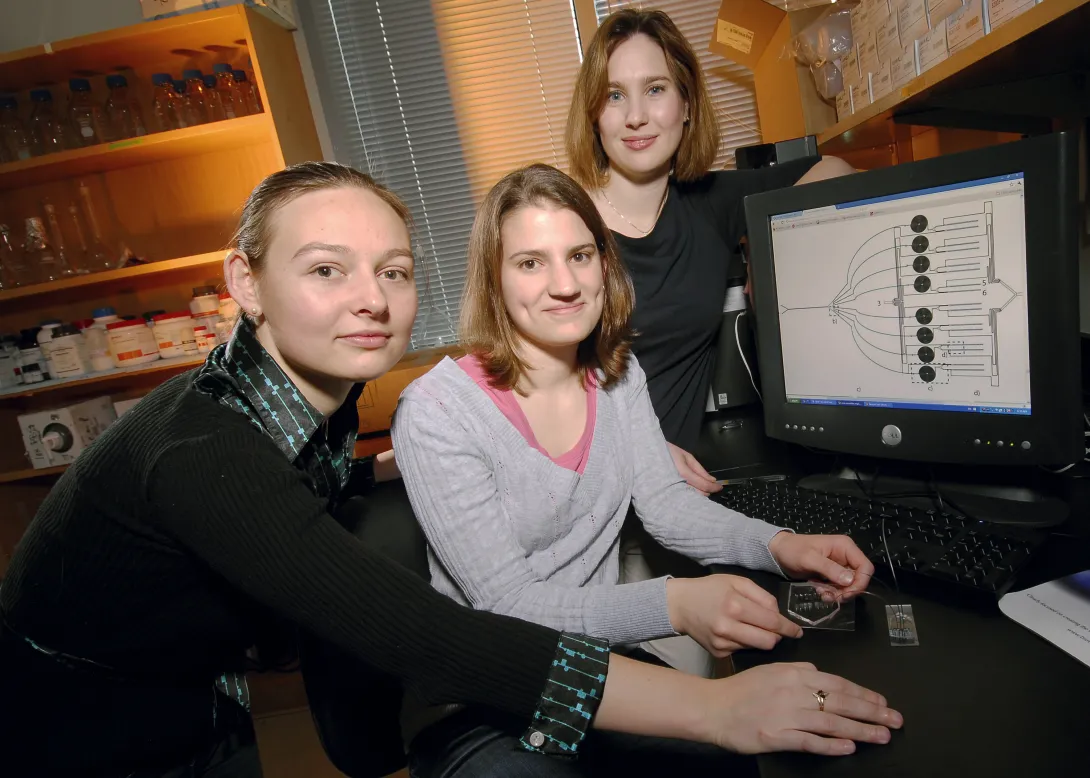
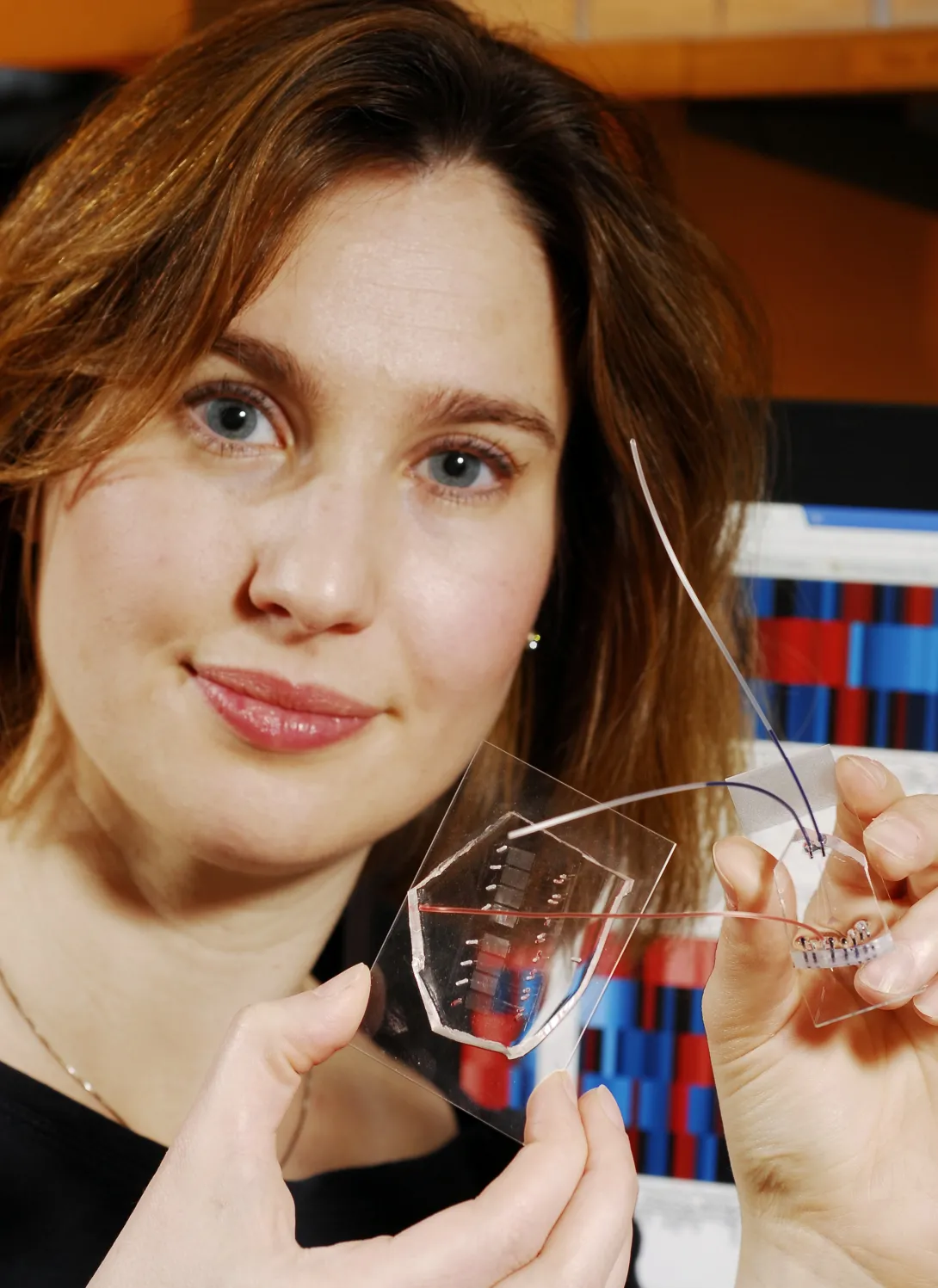
Manipulation of cells by a new microfluidic device may help clinicians improve a promising cancer therapy that harnesses the body's own immune cells to fight such diseases as metastatic melanoma, non-Hodgkin's lymphoma, chronic lymphocytic leukemia and neuroblastoma.
The therapy, known as adoptive T cell transfer, has shown encouraging results in clinical trials. This treatment involves removing disease-fighting immune cells called T cells from a cancer patient, multiplying them in the laboratory and then infusing them back into the patient's body to attack the cancer. The effectiveness of this therapy, however, is limited by the finite lifespan of T cells -- after many divisions, these cells become unresponsive and inactive.
Researchers at Georgia Tech and Emory University have addressed this limitation by developing a microfluidic device for sample handling that allows a statistical model to be generated to evaluate cell responsiveness and accurately predict cell "age" and quality. Being able to assess the age and responsiveness of T cells -- and therefore transfer only young functional cells back into a cancer patient's body -- offers the potential to improve the therapeutic outcome of several cancers.
"The statistical model, enabled by the data generated with the microfluidic device, revealed an optimal combination of extracellular and intracellular proteins that accurately predict T cell age," said Melissa Kemp, an assistant professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. "Knowing this information will help facilitate the clinical development of appropriate T cell expansion and selection protocols."
Details on the microfluidic device and statistical model were published in the March issue of the journal Molecular & Cellular Proteomics. This work was supported by the National Institutes of Health, Georgia Cancer Coalition, and Georgia Tech Integrative Biosystems Institute.
Currently, clinicians measure T cell age by using multiple assays that rely on measurements from large cell populations. The measurements determine if cells are exhibiting functions known to appear at different stages in the life cycle of a T cell.
"Since no one measurement is a perfect predictor, it is advantageous to concurrently sample multiple proteins at different time points, which we can do with our microfluidic device," explained Kemp, who is also a Georgia Cancer Coalition Distinguished Professor. "The wealth of information we get from our device for a small number of cells far exceeds a single measurement from a population the same size by another assay type."
For their study, Kemp, electrical engineering graduate student Catherine Rivet and biomedical engineering undergraduate student Abby Hill analyzed CD8+ T cells from healthy blood donors. They acquired information from 25 static biomarkers and 48 dynamic signaling measurements and found a combination of phenotypic markers and protein signaling dynamics -- including Lck, ERK, CD28 and CD27 -- to be the most useful in predicting cellular age.
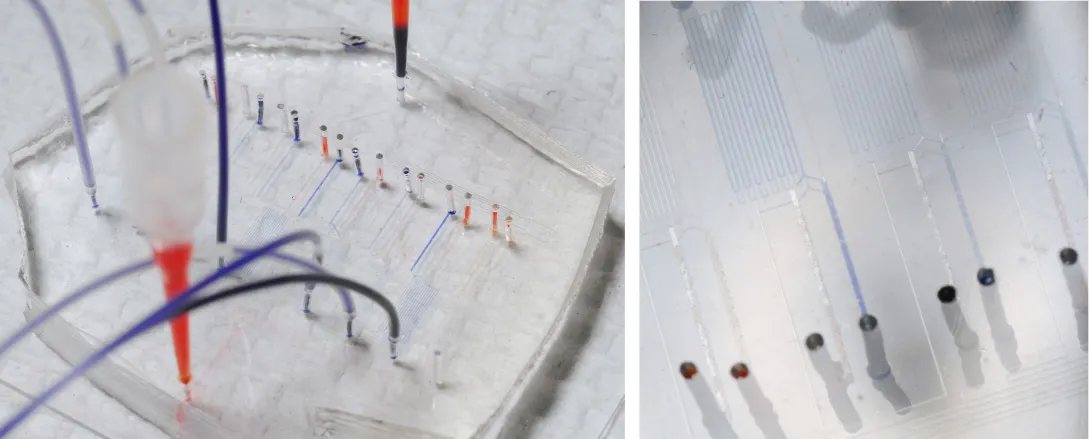
To obtain biomarker and dynamic signaling event measurements, the researchers ran the donor T cells through a microfluidic device designed in collaboration with Hang Lu, an associate professor in the Georgia Tech School of Chemical & Biomolecular Engineering. After stimulating the cells, the device divided them into different channels corresponding to eight different time points, ranging from 30 seconds to seven minutes. Then they were divided again into populations that were chemically treated to halt the biochemical reactions at snapshots in time to build up a picture of the signaling events that occurred as the T cells responded to antigen.
"While donor-to-donor variability is a confounding factor in these types of experiments, the technological platform minimized the experimental data variance and allowed stimulation time to be precisely controlled," said Lu.
With the donor T cell data, the researchers developed a model to assess which biomarkers or dynamical signaling events best predicted the quality of T cell function. The model found the most informative data in predicting cellular age to be the initial changes in signaling dynamics.
"Although a combination of biomarker and dynamic signaling data provided the optimal model, our results suggest that signaling information alone can predict cellular age almost as well as the entire dataset," noted Kemp.
In the future, Kemp plans to use this approach of combining multiple cell-based experiments on a microfluidic chip to integrate single-cell information with population-averaged techniques, such as multiplexed immunoassays or mass spectrometry.
This project is supported in part by the National Institutes of Health (NIH)(Grant No. R21CA134299). The content is solely the responsibility of the principal investigator and does not necessarily represent the official views of the NIH.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364
Feb. 10, 2011
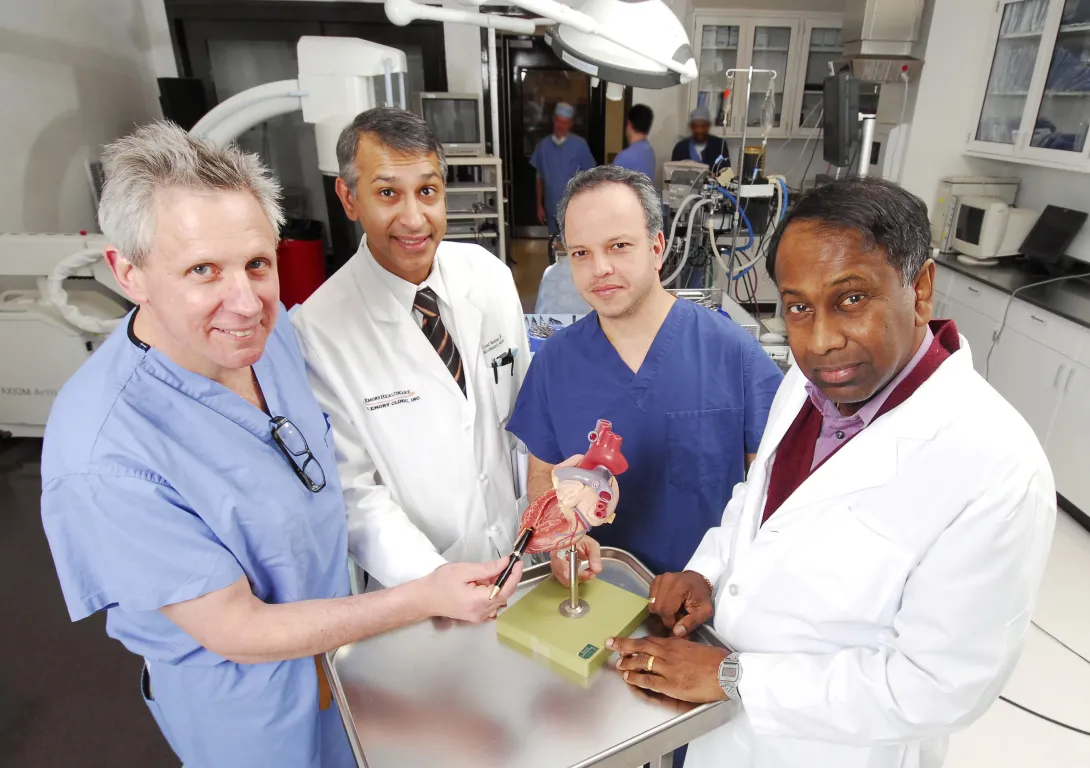
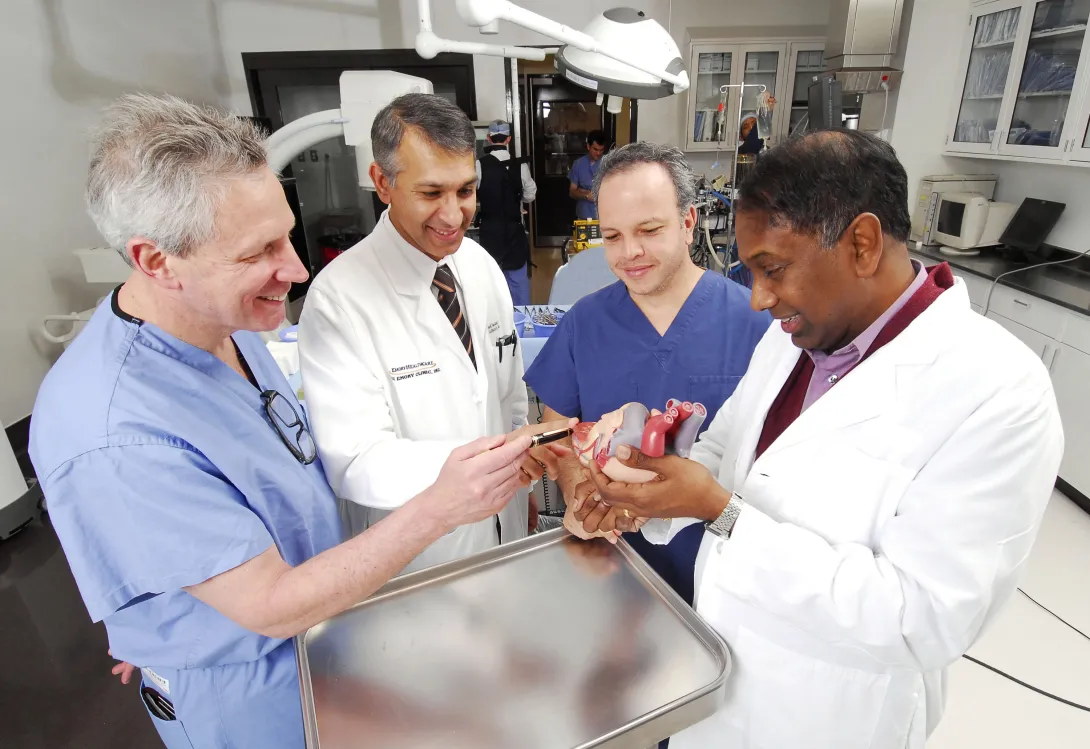
A Georgia Tech and Emory University medical device startup that has developed a system to simplify and standardize the technique for opening and closing the beating heart during cardiac surgery has received a $5.1 million investment.
Apica Cardiovascular has licensed the Georgia Tech/Emory technology and will further develop the system, which will make the transapical access and closure procedure required for delivering therapeutic devices to the heart more routine for all surgeons. The goal is to expand the use of surgery techniques that are less invasive and do not require stopping the heart.
"Our company has leveraged the expertise in cardiovascular technology at Georgia Tech and the clinical experience of surgeons at Emory University to develop a technology that has the potential to revolutionize the delivery of different types of medical devices to the heart, including aortic and mitral valves," said the company's CEO James Greene.
With research and development support from the Coulter Foundation Translational Research Program and the Georgia Research Alliance VentureLab program, the company has already completed a series of pre-clinical studies to test the functionality of the device and its biocompatibility.
The improved heart surgery system consists of a conduit with proprietary technology inside that allows the conduit to be securely attached to the beating heart. Surgeons can then deliver therapeutic devices, such as heart valves or left ventricular assist devices, into the beating heart without loss of blood or exposure to air. Once a therapeutic device has been delivered and surgery is complete, the company's system closes and seals the access site with a biocompatible implant. The closure site can be reopened if necessary.
"By minimizing the incision size to gain access to the beating heart and eliminating the need for conventional sutures, our system improves safety, decreases procedure time and reduces the technical challenges associated with these new minimally invasive procedures," explained Vinod Thourani, an associate professor of surgery and associate director of the Structural Heart Center in Emory University's Division of Cardiothoracic Surgery.
With the new investment from Ireland-based Seroba Kernel Life Sciences and Israel-based TriVentures, the company will continue to conduct research and pre-clinical trials in Atlanta, ultimately leading up to regulatory approval. These efforts will be led by Jorge H. Jimenez, the chief technology officer of the company, which is in the VentureLab process at ATDC, Georgia Tech’s startup company accelerator.
"Our goal is to accelerate and expand the adoption of less-invasive therapeutic procedures to a greater number of surgeons and as a result, many underserved patients will receive needed treatment for valve disease and end-stage heart failure," said Ajit Yoganathan, Regents professor and Wallace H. Coulter Distinguished Faculty Chair in Biomedical Engineering in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University.
The startup will also have an office in Ireland, which will benefit from the strong research collaborations between Georgia Tech, Georgia Tech Ireland and the National University of Ireland, Galway.
"We seek to contribute to and benefit from a global innovation ecosystem in ways that accelerate research results to the market while enhancing economic development opportunities here in Georgia," said Stephen E. Cross, Georgia Tech's executive vice president for research. "Apica Cardiovascular is a perfect example of the synergy between our leading edge work in Atlanta, our Irish translational unit GT Ireland, and our partnership with the National University of Ireland, Galway."
Apica Cardiovascular was founded in 2009 based on technology invented by Jimenez, Thourani, Yoganathan and Thomas Vassiliades, who was an associate professor of cardiothoracic surgery at Emory University at the time. The company was named Emory University's Startup Company of 2010.
About ATDC:
The Advanced Technology Development Center (ATDC) is a startup accelerator that helps technology entrepreneurs in Georgia launch and build successful companies. Founded in 1980, ATDC has graduated more than 120 companies, which together have raised more than a billion dollars in outside financing. In 2010, ATDC was named to Forbes Magazine’s list of the “10 technology incubators that are changing the world.”
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
Jan. 18, 2011
The Tissue Engineering and Regenerative Medicine International Society (TERMIS) will partner with the Parker H. Petit Institute for Bioengineering and Bioscience at Georgia Tech in 2013 by hosting its annual North American Conference in Atlanta.
Two members of the Petit Institute’s faculty have been designated to head the 2013 TERMIS conference. The conference chair will be Robert E. Guldberg, Ph.D., the director of the Petit Institute and professor in mechanical engineering and the program chair will be Todd McDevitt, PhD, associate professor in biomedical engineering and the director of the Stem Cell Engineering Center at Georgia Tech.
“We are honored to be selected and look forward to putting on a great meeting,” Guldberg said. TERMIS brings together an international community to promote discussion of the scientific challenges and therapeutic benefits for the development and application of the tissue engineering and regenerative medicine fields. TERMIS’ mission is to promote worldwide science and technology advancement and education in these fields. It does so through regular worldwide conferences, publishing the Tissue Engineering journal that it endorses and providing quarterly newsletters and other communications for its members.
“Bob and I are dedicated to creating a dynamic program that will honor the meetings of the past as well as introduce some new elements,” McDevitt remarked. TERMIS has been evolving over the last decade. Its roots began in 2001 as an annual workshop called “Tissue Growth Engineering” that was organized by the Pittsburgh Tissue Engineering Initiative. In 2004, this small workshop evolved into the larger, national meeting called Regenerate. In 2006, the Regenerate World Congress was held in Pittsburgh where the meeting had grown significantly and had a large international following. By the following year, the society was rebranded into TERMIS to encompass its international presence. The society has continued to grow and now has chapters in Europe and Asia. TERMIS is open to anyone engaged in research in the tissue engineering or regenerative medicine arenas.
The 2011 TERMIS North American conference was held in Houston, Texas and in 2012 the entire society will come together for the TERMIS World Congress in Vienna, Austria.
News Contact
Megan McDevitt
IBB
Contact Megan McDevitt
404-385-7001
Sep. 23, 2010
Phillip Santangelo, assistant professor in the Coulter Department, has received an R01 NIH/National Institute for General Medicine Sciences award to develop single molecule sensitive probes for the study of virus replication, assembly and budding. The $1.48 million project will focus on the human respiratory syncytial (hRSV) virus. hRSV is recognized as the most important viral agent of serious pediatric respiratory tract disease. Worldwide, acute respiratory tract disease is the leading cause of mortality due to infectious disease, and hRSV remains one of the pathogens deemed most important for vaccine and antiviral development. He will collaborate with James E. Crowe, Jr., MD, The Departments of Microbiology and Immunology, and Pediatrics and The Vanderbilt Vaccine Center; Vanderbilt University Medical Center for the 5-year study.
News Contact
Adrianne ProellerWallace H. Coulter Dept. of Biomedical Engineering
Contact Adrianne Proeller
Sep. 24, 2010
Atlanta (September 24, 2010) — Assistant Professor Francesca Storici (Biology) has been awarded a research grant by the National Science Foundation (NSF) for a 3 year project focusing on “Mechanisms of RNA/DNA hybrid stability and of information flow from RNA to DNA in yeast cells". The goal of this research is to understand the mechanisms by which RNA can directly transfer information to the DNA of cells. The main objectives are: 1) to identify the main protein factors cleaving the RNA tract in an RNA/DNA hybrid during RNA-driven DNA repair and DNA modification and to characterize their in vivo functions, and 2) to reveal the role of DNA repair mechanisms in the removal of RNA embedded into DNA. This project addresses challenging questions in molecular biology: How likely is information flow from RNA to DNA in cells? How well is RNA tolerated in DNA? What are the consequences of RNA-driven modifications in cells? The study will be done using newly developed systems in the yeast Saccharomyces cerevisiae, which will be exploited to perform molecular and cellular biology experiments to identify and characterize the molecular mechanisms of RNA-driven DNA repair and editing.
Related Links
Pagination
- Previous page
- Page 46
- Next page