Jun. 14, 2023
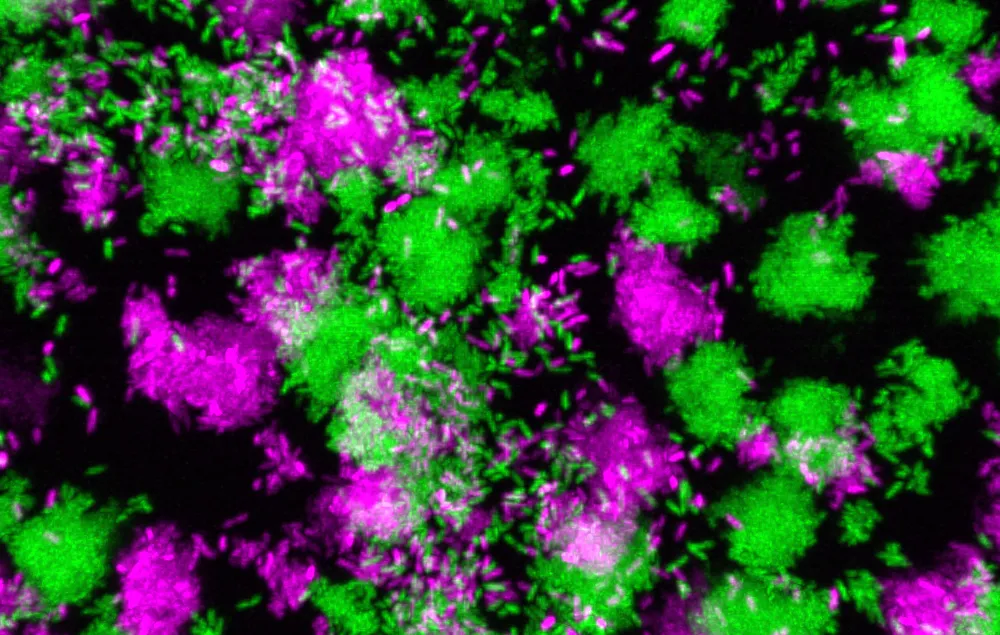
People with weakened immune systems are at constant risk of infection. Pseudomonas aeruginosa, a common environmental bacterium, can colonize different body parts, such as the lungs, leading to persistent, chronic infections that can last a lifetime – a common occurrence in people with cystic fibrosis.
But the bacteria can sometimes change their behavior and enter the bloodstream, causing chronic localized infections to become acute and potentially fatal. Despite decades of studying the transition in lab environments, how and why the switch happens in humans has remained unknown.
However, researchers at the Georgia Institute of Technology have identified the major mechanism behind the transition between chronic and acute P. aeruginosa infections. Marvin Whiteley – professor in the School of Biological Sciences and Bennie H. and Nelson D. Abell Chair in Molecular and Cellular Biology – and Pengbo Cao, a postdoctoral researcher in Whiteley’s lab, discovered a gene that drives the switch. By measuring bacterial gene expression in human tissue samples, the researchers identified a biomarker for the transition.
Their research findings, published in Nature, can inform the development of future treatments for life-threatening acute infections.
According to Whiteley and Cao, bacteria, like animals, are versatile and behave differently depending on their environment. A person with a chronic infection might be fine one day, but environmental changes in the body can cause bacteria to change their behavior. This can lead to acute infection, and a person could develop sepsis that requires immediate treatment.
“For years, people have been studying these bacteria in well-controlled lab environments, even though the lab is a place most microbes have never seen,” said Whiteley. “Our study took a novel approach to look directly into the bacterium’s behavior in the human host.”
The researchers chose to look at human tissue samples of chronic bacterial lung and wound infections. Using genetic sequencing technologies, Whiteley and Cao measured the levels of all types of mRNA present in the bacteria. The mRNAs encode the proteins that do all the work in a cell, so by measuring a bacterium’s mRNA level, one can infer the bacterium’s behavior.
While P. aeruginosa has roughly 6,000 genes, Whiteley and Cao found that one gene in particular – known as PA1414 – was more highly expressed in human tissue samples than all the other thousands of genes combined. The levels were so high that, at first, Cao and Whiteley thought the amount of PA1414 mRNA might be an artifact – a glitch associated with the sequencing methods.
“This particular gene is not expressed in the standard lab environment very much, so it was striking to see these levels,” Cao said. “And at this point, the function of the gene was unknown.”
The researchers also found that low oxygen drives the high expression of the gene. This is a common environmental characteristic of bacterial infections, as bacteria frequently encounter oxygen deprivation during chronic infections. Further tests showed that the gene also regulates bacterial respiration under low oxygen conditions.
Interestingly, the researchers found that rather than encoding a protein, the gene encodes a small RNA that plays a vital role in bacterial respiration. They named the small RNA SicX (sRNA inducer of chronic infection X).
The researchers then tested the functions of the gene in different animal infection models. They observed that when SicX wasn’t present, the bacteria easily disseminated from chronic infections throughout the body, causing systemic infection. The comparison allowed the researchers to determine that the gene is important for promoting chronic localized infection. Moreover, researchers also showed that the expression of SicX immediately decreased during the transition from chronic to acute infection, suggesting SicX potentially serves as a biomarker for the chronic-to-acute switch.
“In other words, without the small RNA, the bacteria become restless and go looking for oxygen, because they need to breathe like we need to breathe,” Whiteley said. “That need causes the bacteria to enter the bloodstream. Now, we know that oxygen levels are regulating this transition.”
Having a better indication for when an infection might enter the bloodstream would be a paradigm shift for treatments.
“If you can predict when an acute infection will occur, a patient could take a diagnostic test at home to determine if and when they may need to get treatment – before the infection becomes life-threatening,” Whiteley said.
The study provides answers to the long-standing questions about how and why chronic infections become acute. The researchers’ findings also open opportunities to develop therapeutics that target this specific molecular behavior associated with P. aeruginosa infections.
“The chronic Pseudomonas infection is usually highly resistant to first-line antibiotics,” Cao said. “By targeting this small RNA, we could potentially change the lifestyle of the bacteria to make it more susceptible to antibiotic treatments and achieve greater clearance of these dangerous infections.”
Marvin Whiteley is also an Eminent Scholar with the Georgia Research Alliance.
Citation: Cao, P., Fleming, D., Moustafa, D.A., et al. A Pseudomonas aeruginosa small RNA regulates chronic and acute infection. Nature 618, 358–364 (2023).
DOI: https://doi.org/10.1038/s41586-023-06111-7
Funding: NIH grants R21AI154220, R21AI137462, and R21AI147178; Cystic Fibrosis Foundation grants WHITEL20A0 and WHITEL22G0; Cystic Fibrosis Trust Foundation grant SRC017; and Cystic Fibrosis Postdoctoral Fellowships CAO20F0 and DOLAN20F0
News Contact
Catherine Barzler, Senior Research Writer/Editor
May. 18, 2023
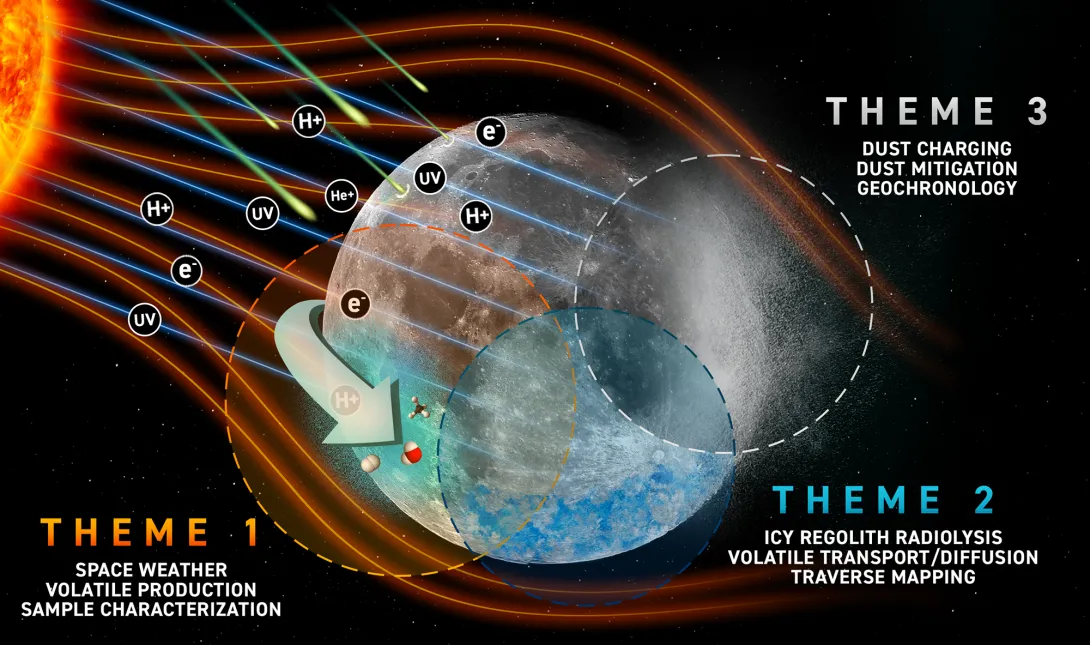
Georgia Tech researchers have been selected by NASA to lead a $7.5 million center that will study the lunar environment and the generation and properties of volatiles and dust. The Center for Lunar Environment and Volatile Exploration Research (CLEVER) will be led by Thomas Orlando, professor in the School of Chemistry and Biochemistry.
CLEVER is the successor to Orlando’s pioneering REVEALS (Radiation Effects on Volatiles and Exploration of Asteroids and Lunar Surfaces) center, and both are part of NASA’s Solar System Exploration Research Virtual Institute (SSERVI) program.
REVEALS and CLEVER look ahead to the return of humans to the moon for sustained periods — a key part of NASA’s plan for space exploration in the coming decade. Volatiles such as water, molecular oxygen, methane, and hydrogen are crucial to supporting human activity on the moon. Dust is also important since the space-weathered particles can pose health effects to astronauts and hazards to the technology and hardware.
The interdisciplinary group of researchers supported by CLEVER will study how the solar wind and micrometeorites produce volatiles, research how ice and dust behave in the lunar environment, develop new materials to deal with potential dust buildup, and invent new analysis tools to support the upcoming crewed missions of the Artemis program.
“The resources and knowledge that CLEVER will produce will be useful for the sustainable presence of humans on the moon,” Orlando says. “We have the correct mix of fundamental science and exploration — real, fundamental, ground-truth measurements; very good theory/modeling; and engineering — an easy mix with Georgia Tech and outside partners.”
Orlando adds that CLEVER adopts a unique perspective on the challenges of understanding how to operate on Earth’s moon. “The atomic and molecular view of processes with angstrom distances and femtosecond time scales can help unravel what is happening on planetary spatial scales and geological time frames,” he says. “We can also translate our knowledge into materials, devices, and technology pretty quickly, and this is necessary if we want to help the Artemis astronauts.”
CLEVER includes investigators from Georgia Tech, University of Georgia, the Florida Space Institute, University of Hawaii, Auburn University, Space Sciences Institute, the Johns Hopkins University Applied Physics Laboratory, Lawrence Berkeley National Laboratory, NASA Ames, NASA Kennedy Space Center, and partners in Italy and Germany. In addition to pursuing a blend of fundamental science and mission support, CLEVER will also emphasize the research and career development of students and young investigators, another important goal of the SSERVI system.
Learn more about the Center on Lunar Research and Exploration by visiting their website.
Writer: M.G. Finn
Art: Brice Zimmerman
News Contact
Catherine Barzler, Senior Research Writer/Editor
May. 11, 2023
A team of researchers from Georgia Tech, Emory University, Morehouse School of Medicine, University of Georgia, the Center for Global Health Innovation, and the Technical College System of Georgia has been awarded $1 million over the course of two years from the U.S. National Science Foundation's Regional Innovation Engines, or NSF Engines, program. They are among the more than 40 unique teams to receive one of the first-ever NSF Engines Development Awards, which aim to help partners collaborate to create economic, societal, and technological opportunities for their regions.
The team, “Advancing Health Equity and Diagnostic Technologies (GA) Development,” will use the award to support key institutional, corporate, government, education, and community partners to create an innovative ecosystem that will inspire, develop, and translate affordable and widely available point-of-care (POC) medical technologies to advance health equity throughout the southeast.
“The Southeastern U.S. has the lowest life expectancy in the nation, and there are significant health disparities along economic, educational, racial, and geographic divisions,” said Wilbur Lam, Georgia Tech professor, principal investigator, and innovation lead. “The team will work to build an ecosystem of partners to drive use-inspired research and technology translation in the area of POC diagnostics and wearables with strong community engagement to help address these areas and advance health equity.”
The NSF Engines program is a transformational investment for the nation, ensuring the U.S. remains in the vanguard of competitiveness for decades to come.
"These NSF Engines Development Awards lay the foundation for emerging hubs of innovation and potential future NSF Engines," said NSF Director Sethuraman Panchanathan. "These awardees are part of the fabric of NSF's vision to create opportunities everywhere and enable innovation anywhere. They will build robust regional partnerships rooted in scientific and technological innovation in every part of our nation. Through these planning awards, NSF is seeding the future for in-place innovation in communities and to grow their regional economies through research and partnerships. This will unleash ideas, talent, pathways and resources to create vibrant innovation ecosystems all across our nation."
Led by Lam, the team aims to build an ecosystem to drive use-inspired research and technology translation for health equity and leverage relationships with underserved Georgia communities to inspire a technology roadmap and adopt new technologies. An annual event and comprehensive roadmap will drive sustainable technology translation, workforce development, and systemic education.
The awardees span a broad range of states and regions, reaching geographic areas that have not fully benefited from the technology boom of the past decades. These NSF Engines Development Awards will help organizations create connections and develop their local innovation ecosystems within two years to prepare strong proposals for becoming future NSF Engines, which will each have the opportunity to receive up to $160 million.
Launched by NSF's new Directorate for Technology, Innovation and Partnerships and authorized by the "CHIPS and Science Act of 2022," the NSF Engines program uniquely harnesses the nation's science and technology research and development enterprise and regional-level resources. NSF Engines aspire to catalyze robust partnerships to positively impact regional economies, accelerate technology development, address societal challenges, advance national competitiveness and create local, high-wage jobs.
View a map of the NSF Engines Development Awards. More information can be found on the NSF Engines program website.
NSF MEDIA REQUESTS: media@nsf.gov
GEORGIA TECH MEDIA REQUESTS: georgia.parmelee@gatech.edu
May. 10, 2023
The world would look very different without multicellular organisms – take away the plants, animals, fungi, and seaweed, and Earth starts to look like a wetter, greener version of Mars. But precisely how multicellular organisms evolved from single-celled ancestors remains poorly understood. The transition happened hundreds of millions of years ago, and early multicellular species are largely lost to extinction.
To investigate how multicellular life evolves from scratch, researchers from the Georgia Institute of Technology decided to take evolution into their own hands. Led by William Ratcliff, associate professor in the School of Biological Sciences and director of the Interdisciplinary Graduate Program in Quantitative Biosciences, a team of researchers has initiated the first long-term evolution experiment aimed at evolving new kinds of multicellular organisms from single-celled ancestors in the lab.
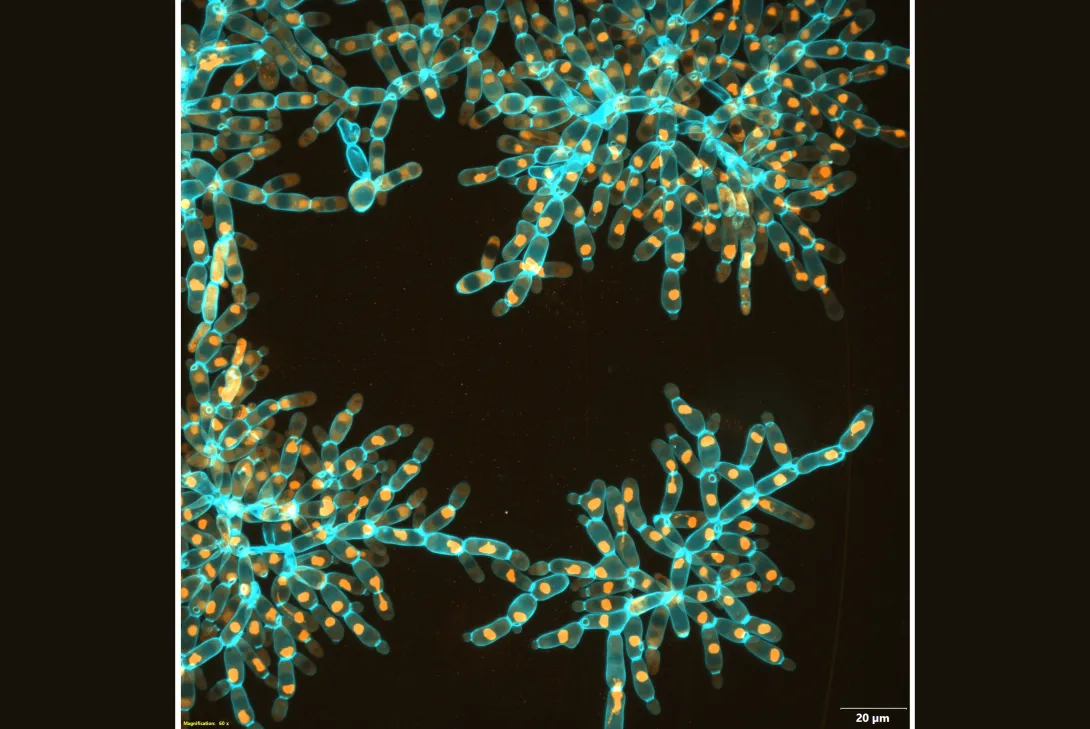
Over 3,000 generations of laboratory evolution, the researchers watched as their model organism, “snowflake yeast,” began to adapt as multicellular individuals. In research published in Nature, the team shows how snowflake yeast evolved to be physically stronger and more than 20,000 times larger than their ancestor. This type of biophysical evolution is a pre-requisite for the kind of large multicellular life that can be seen with the naked eye. Their study is the first major report on the ongoing Multicellularity Long-Term Evolution Experiment (MuLTEE), which the team hopes to run for decades.
“Conceptually, what we want to understand is how simple groups of cells evolve into organisms, with specialization, coordinated growth, emergent multicellular behaviors, and life cycles – the stuff that differentiates a pile of pond scum from an organism that is capable of sustained evolution,” Ratcliff said. “Understanding that process is a major goal of our field.”
The Multicellularity Long-Term Evolution Experiment
Ozan Bozdag, a research scientist and former postdoctoral researcher in Ratcliff’s group and first author on the paper, initiated the MuLTEE in 2018, starting with single-celled snowflake yeast. Bozdag grew the yeast in shaking incubators and each day selected for both faster growth and larger group size.
The team selected on organism size because all multicellular lineages started out small and simple, and many evolved to be larger and more robust over time. The ability to grow large, tough bodies is thought to play a role in increasing complexity, as it requires new biophysical innovations. However, this hypothesis had never been directly tested in the lab.
Over about 3,000 generations of evolution, their yeast evolved to form groups that were more than 20,000 times larger than their ancestor. They went from being invisible to the naked eye to the size of fruit flies, containing over half a million cells. The individual snowflake yeast evolved novel material properties: while they started off weaker than gelatin, they evolved to be as strong and tough as wood.
New Biophysical Adaptations
In investigating how the snowflake yeast adapted to become larger, the researchers observed that the yeast cells themselves became elongated, reducing the density of cells packed into the group. This cell elongation slowed down the accumulation of cell-to-cell stress that would normally cause the clusters to fracture, allowing the groups to get larger. But this fact alone should have only resulted in small increases in size and multicellular toughness.
To uncover the precise biophysical mechanisms that allowed growth to macroscopic size, the researchers needed to look inside the yeast clusters to see how the cells interacted physically. Normal light microscopes were unable to penetrate the large, densely packed groups, so the researchers used a scanning electron microscope to image thousands of ultrathin slices of the yeast, which gave them their internal structure.
“We discovered that there was a totally new physical mechanism that allowed the groups to grow to this very, very large size,” Bozdag said. “The branches of the yeast had become entangled – the cluster cells evolved vine-like behavior, wrapping around each other and strengthening the entire structure.”
By simply selecting on organismal size, the researchers figured out how to leverage the biomechanical mechanism of entanglement, which ended up making the yeast about 10,000 times tougher as a material.
“Entanglement has previously been studied in totally different systems, mostly in polymers,” said Peter Yunker, associate professor in the School of Physics and a co-author on the paper. “But here we’re seeing entanglement through an entirely different mechanism — the growth of cells rather than just through their movement.”
Observing the entanglement was a turning point in the researchers’ understanding of how simple multicellular groups evolve. As a brand-new multicellular organism, snowflake yeast lacks the sophisticated developmental mechanisms that characterize modern multicellular organisms. But after just 3,000 generations of laboratory evolution, the yeast figured out how to drive and co-opt cellular entanglement as a developmental mechanism.
Preliminary investigations of other multicellular fungi show that they also form highly entangled multicellular bodies, suggesting that entanglement is a widespread and important multicellular trait in this branch of multicellular life.
“I’m really excited to have a model system where we can evolve early multicellular life over thousands of generations, harnessing the awesome power of modern science,” Ratcliff said. “In principle, we can understand everything that is happening, from the evolutionary cell biology to the biophysical traits which are directly under selection.”
For a long time, humans have worked with biology to evolve new things – from the corn we eat to domesticated dogs, chickens, and show pigeons. According to Ratcliff, what their team is doing is not so different.
“By putting our finger on the scale of a single-celled organism’s evolution, we can figure out how they evolved into progressively more complex and integrated multicellular organisms, and can study that process along the way,” he said. “We hope that this is just the first chapter in a long story of multicellular discovery as we continue to evolve snowflake yeast in the MuLTEE.”
Citation: Bozdag GO, Zamani-Dahaj SA, Day TC, Kahn PC, Burnetti AJ, Lac DT, Tong K, Conlin PL, Balwani AH, Dyer EL, Yunker PJ. De novo evolution of macroscopic multicellularity. Nature, 2023.
DOI: 10.1038/s41586-023-06052-1
News Contact
Catherine Barzler, Senior Research Writer/Editor
Apr. 28, 2023
One of the National Academy of Engineering’s Grand Challenges for Engineering is to engineer better medicines. To help address this challenge, W. Hong Yeo leads the Materials for Biomedical Systems research initiative for the Georgia Tech Institute for Materials (IMat). The goal of the initiative is to enhance human health via multidisciplinary materials research.
“The existing healthcare challenges are so complicated and demanding, so the collaboration between academia, industry, and national labs is imperative, and synergistic multidisciplinary research is required,” explained Yeo, who is also an associate professor and Woodruff Faculty Fellow in the Woodruff School of Mechanical Engineering and holds a courtesy appointment in the Coulter Department of Biomedical Engineering.
To further this initiative, Yeo and Emory University’s Young Jang organized the Materials for Biomedical Systems (MBS) Day at Georgia Tech. The workshop was held on March 30 at the Georgia Tech Global Learning Center and attracted researchers and industry representatives from a variety of disciplines.
The focus of the morning session was on soft materials and biomaterials for medical systems. It began with a talk on Organogels x EGaIn for Soft & Self-Healing Bioelectronics from Carmel Majidi, a professor of mechanical engineering at Carnegie Mellon University. Additional speakers in the morning session included ProgenaCare Global’s Allison Ramey-Ward, Seoul National University’s Young Bin Choy, and Korea Advanced Institute of Science & Technology’s Jae-Woong Jeong. The morning concluded with a panel discussion, regarding the translation of biomaterials technologies to system developments and commercialization, moderated by the University of Pittsburgh’s Youngjae Chun.
The afternoon session of the day was focused on stem cells and regenerative medicine. It began with a talk on Bioengineered Hydrogels for Regenerative Medicine from Andrés García, executive director of the Petit Institute for Bioengineering and Bioscience (IBB) and Regents’ Professor at Georgia Tech. Additional speakers in this session included Sung-Jin Park from Emory/Georgia Tech, William Hynes from Lawrence Livermore National Laboratory, Ki Dong Park from Ajou University, Ho-Wook Jun from the University of Alabama at Birmingham, and Johnna Temenoff from Emory University/Georgia Tech. The session concluded with a panel discussion moderated by Johnny Lam from the Food and Drug Administration.
“I am so thankful for all of the participants, sponsors, and organizers who made such an amazing workshop that generated innovative ideas and new collaboration opportunities from across the field,” said Yeo. “We also discussed immediate commercialization paths and regulatory importance in developing biomaterials and medical systems. We will continue offering networking and research-sharing opportunities to facilitate knowledge exchange through this MBS initiative.”
After the workshop, multiple students participated in a poster contest to showcase their research in biomaterials and medical systems and network with attendees. MBS Day was co-sponsored by IMat and IBB.
News Contact
Laurie Haigh
Research Communications
Apr. 06, 2023
Professor Omer Inan is set to take the stage at the upcoming TEDxAtlanta 2023: We Rise event on May 19.
As the Linda J. and Mark C. Smith Chair in bioscience and bioengineering in Tech’s School of Electrical and Computer Engineering (ECE), Inan designs clinically relevant medical devices and systems and translates them from the lab to patient care applications. In his talk, Inan will be discussing his groundbreaking research on wearable healthcare technologies and the potential they hold for revolutionizing the field.
Inan is a member of the prestigious Medical and Biological Engineering (AIMBE) College of Fellows (elected in 2022) for his “outstanding contributions to the non-invasive assessment of the mechanical aspects of cardiovascular health and performance using wearable devices.” Additional achievements include an Academy Award for Technical Achievement from The Academy of Motion Picture Arts and Sciences (The Oscars, 2021), the Georgia Power Professor of Excellence for the College of Engineering (2019), and the National Science Foundation Faculty Early Career Development Program award (NSF CAREER, 2018).
TEDxAtlanta 2023: WE RISE brings together an impressive group of participants from diverse backgrounds, experiences, and perspectives. The speakers include entrepreneurs, activists, educators, artists, scientists, and many other changemakers who have risen above challenges to make a positive impact on the world.
The event's participants will share their stories and insights on how they have overcome adversity, embraced innovation, and challenged the status quo to make a difference in their communities and beyond. Through their talks, they will inspire and empower attendees to rise above their own challenges and take action towards creating a better future for all.
TEDxAtlanta 2023: WE RISE will take place on Friday, May 19 from 9 a.m. – 6:30 p.m. at the Rialto Center for the Arts (80 Forsyth Street Northwest Atlanta, GA 30303). Learn more and purchase tickets at tedxatlanta.com.
News Contact
Dan Watson
Mar. 31, 2023
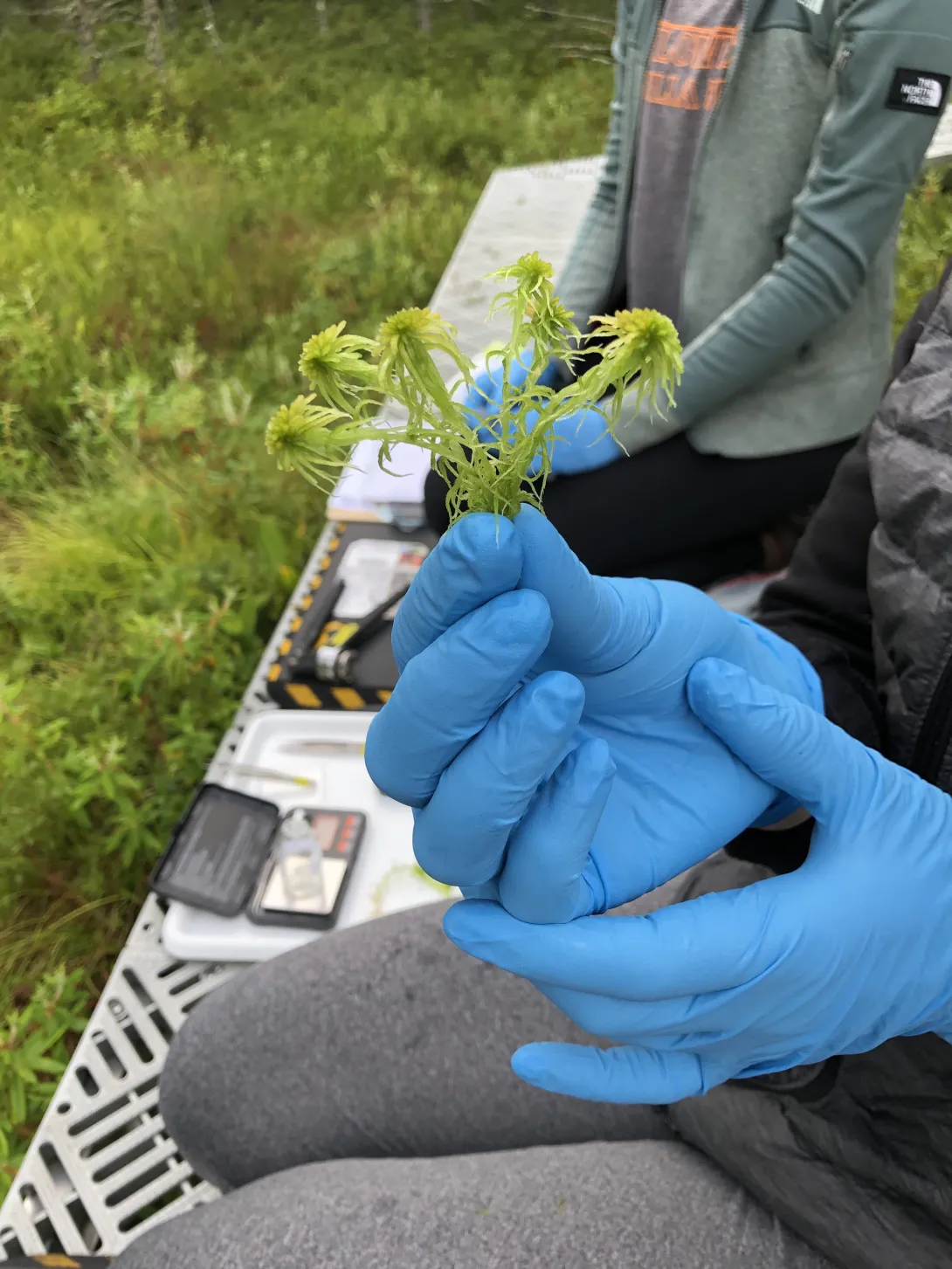
If you’re an avid gardener, you may have considered peat moss — decomposed Sphagnum moss that helps retain moisture in soil — to enhance your home soil mixture. And while the potting medium can help plants thrive, it’s also a key component of peatlands: wetlands characterized by a thick layer of water-saturated, carbon-rich peat beneath living Sphagnum moss, trees, and other plant life.
These ecosystems cover just 3% of Earth’s land area, but “peatlands store over one-third of all soil carbon on the planet,” explains Joel Kostka, professor and associate chair of Research in the School of Biological Sciences at Georgia Tech.
This carbon storage is supported in large part by microbes. Two microbial processes in particular — nitrogen fixation and methane oxidation — strike a delicate balance, working together to give Sphagnum mosses access to critical nutrients in nutrient-depleted peatlands.
The coupling of these two processes is often referred to as the “missing link” of nutrient cycling in peatlands. Yet, how these processes will respond to changing climates along northern latitudes is unclear.
“There are tropical peatlands — but the majority of peatlands are in northern environments.” notes Caitlin Petro, a research scientist who works with Kostka in Biological Sciences at Tech. “And those are going to be hit harder by climate change.”
Kostka and Petro recently led a collaborative study to investigate how this critical type of ecosystem (and the “missing link” of microbial processes that support it) may react to the increased temperature and carbon dioxide levels predicted to come with climate change. The team, which also includes researchers from the Oak Ridge National Laboratory (ORNL), Florida State University, and the University of Tennessee, Knoxville, just published their work in the scientific journal Global Change Biology.
By testing the effects of increasing temperature and carbon dioxide on the growth of Sphagnum moss, its associated microbiome, and overall ecosystem health, Kostka and Petro say computational models will be better equipped to predict the effects of climate change.
“Down the road,” Kostka added, “we hope the results can be used by environmental managers and governments to adaptively manage or geoengineer peatlands to thrive in a warmer world.”
Raising the heat
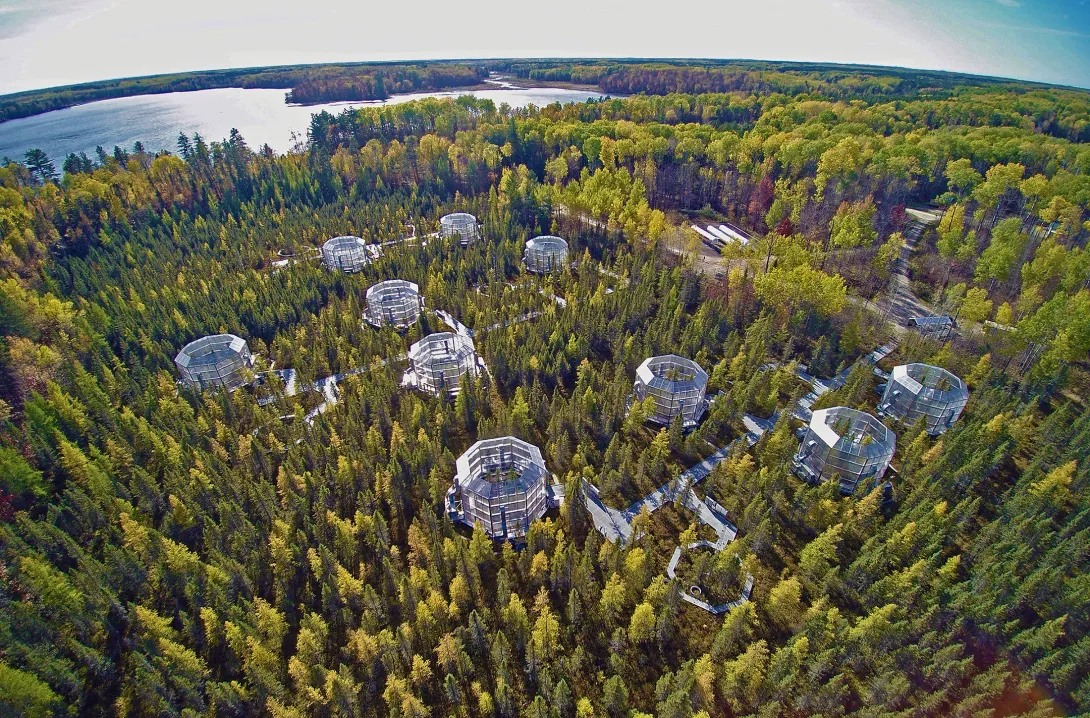
To see how northern peatlands will react to climate change, the team, which also included School of Earth and Atmospheric Sciences Associate Professor Jennifer Glass, turned to the ORNL Spruce and Peatland Responses Under Changing Environments (SPRUCE) experiment — a unique field lab in northern Minnesota where the team warms peat bogs and experimentally changes the amount of carbon dioxide in the atmosphere.
Starting in 2016, the team exposed different parts of SPRUCE’s experimental peatlands to a gradient of higher temperatures ranging from an increase of 0°C to 9°C, capturing the Intergovernmental Panel on Climate Change models’ predicted 4°C to 6°C increase in northern regions by 2100.
The moss’s reaction was significant. Although nearly 100% of the bog’s surface was covered in moss at the beginning of the experiment, moss coverage dropped with each increase in temperature, plummeting to less than 15% in the warmest conditions.
Critically, the two microbial processes that had previously been consistently linked fell out of sync at higher temperatures.
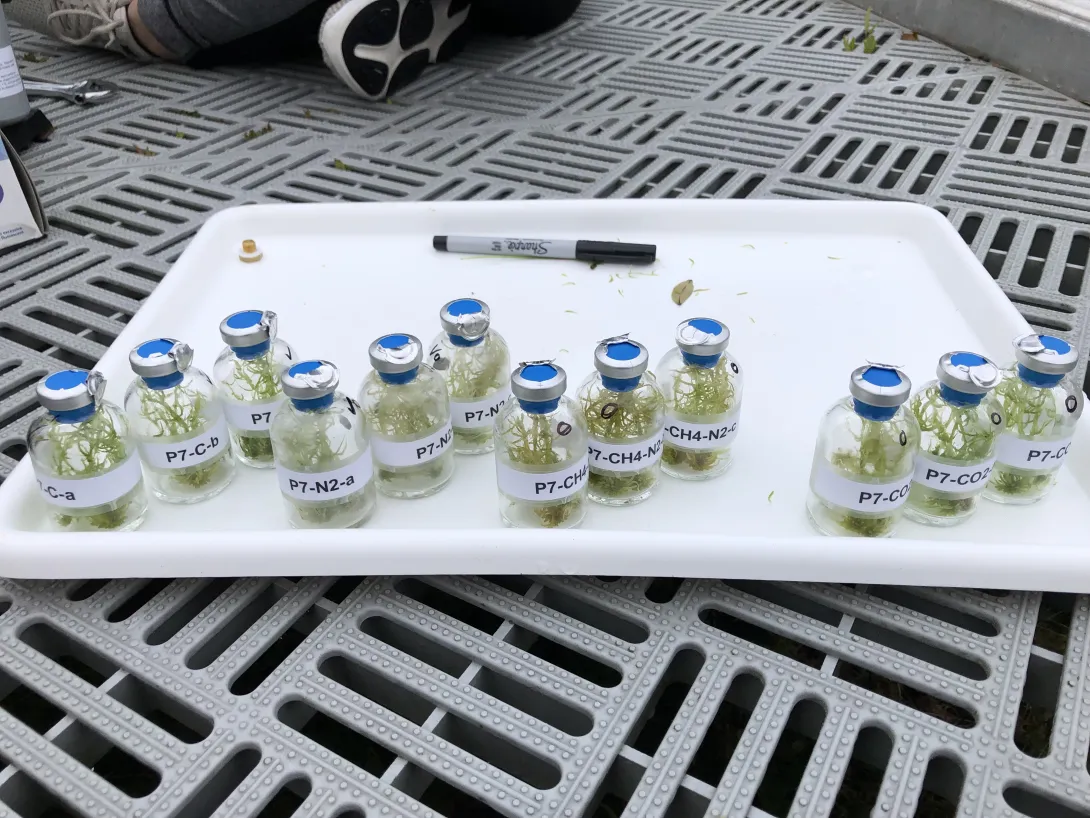
“Peatlands are extremely nutrient-poor and microbial nitrogen fixation represents a major nitrogen input to the ecosystem,” Kostka explained. Fixing nitrogen is the process of turning atmospheric nitrogen into an organic compound that the moss can use for photosynthesis, while methane oxidation allows the moss to use methane released from decomposing peat as energy. “Methane oxidation acts to fuel nitrogen fixation while scavenging a really important greenhouse gas before it is released to the atmosphere. This study shows that these two processes, which are catalyzed by the Sphagnum microbiome, become disconnected as the moss dies.”
“These processes occurring together are really important for the community,” Petro explained. Yet many microbes that are able to both fix nitrogen and oxidize methane were absent in the mosses collected from higher temperature enclosures. And while elevated carbon dioxide levels appeared to offset some of the changes in nitrogen cycling caused by warming, the decoupling of these processes remained.
“These treatments are altering a fairly well-defined and consistent plant microbiome that we find in many different environments, and that has this consistent function,” Petro explained. “It's like a complete functional shift in the community.”
Though it’s not clear which of these changes — the moss dying or the altered microbial activity — is driving the other, it is clear that with warmer temperatures and higher carbon dioxide levels comes a cascade of unpredictable outcomes for peat bogs.
“In addition to the direct effects of climate warming on ecosystem function,” Petro adds, “it will also introduce all of these off-shooting effects that will impact peatlands in ways that we didn't predict before.”
This work was supported by the National Science Foundation (DEB grant no. 1754756). The SPRUCE project is supported by the U.S. Department of Energy's Office of Science, Biological, and Environmental Research (DOE BER) and the USDA Forest Service.
DOI: https://doi.org/10.1111/gcb.16651
Citation: Petro, C., et al. Climate drivers alter nitrogen availability in surface peat and decouple N2 fixation from CH4 oxidation in the Sphagnum moss microbiome. Global Change Biology. (2023).
Aerial Photo: Hanson, P.J., M.B. Krassovski, and L.A. Hook. 2020. SPRUCE S1 Bog and SPRUCE Experiment Aerial Photographs. Oak Ridge National Laboratory, TES SFA, U.S. Department of Energy, Oak Ridge, Tennessee, U.S.A. https://doi.org/10.3334/CDIAC/spruce.012 (UAV image number 0050 collected on October 4, 2020).
News Contact
Writer: Audra Davidson
Communications Officer II, College of Sciences
Editor: Jess Hunt-Ralston
Director of Communications, College of Sciences
Mar. 02, 2023
Ryan Lawler realized early on in her academic career that a scientist with a great idea can potentially change the world.
“But I didn’t realize the role that real estate can play in that,” said Lawler, general manager of BioSpark Labs – the collaborative, shared laboratory environment taking shape at Science Square at Georgia Tech.
Sitting adjacent to the Tech campus and formerly known as Technology Enterprise Park, Science Square is being reactivated and positioned as a life sciences research destination. The 18-acre site is abuzz with new construction, as an urban mixed-use development rises from the property.
Meanwhile, positioned literally on the ground floor of all this activity is BioSpark Labs, located in a former warehouse, fortuitously adjacent to the Global Center for Medical Innovation. It’s one of the newer best-kept secrets in the Georgia Tech research community.
BioSpark exists because the Georgia Tech Real Estate Office, led by Associate Vice President Tony Zivalich, recognized the need of this kind of lab space. Zivalich and his team have overseen the ideation, design, and funding of the facility, partnering with Georgia Advanced Technology Ventures, as well as the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University, and the core facilities of the Petit Institute for Bioengineering and Bioscience.
“We are in the middle of a growing life sciences ecosystem, part of a larger vision in biotech research,” said Lawler, who was hired on to manage the space, bringing to the job a wealth of experience as a former research scientist and lab manager with a background in molecular and synthetic biology.
Researchers’ Advocate
BioSpark was designed to be a launch pad for high-potential entrepreneurs. It provides a fully equipped and professionally operated wet lab, in addition to a clean room, meeting and office space, to its current roster of clients, five life sciences and biotech startup, a number certain to increase – because BioSpark is undergoing a dramatic expansion that will include 11 more labs (shared and private space), an autoclave room, equipment and storage rooms.
“We want to provide the necessary services and support that an early-stage company needs to begin lab operations on day one,” said Lawler, who has put together a facility with $1.7 million in lab equipment. “I understand our clients’ perspective, I understand researchers and their experiments, and their needs, because I have first-hand proficiency in that world. So, I can advocate on their behalf.”
CO2 incubators, a spectrophotometer, a biosafety cabinet, a fume hood, a -80° freezer, an inverted microscope, and the autoclave are among the wide range of apparatus. Plus, a virtual treasure trove of equipment is available to BioSpark clients off-site through the Core Facilities of the Petit Institute for Bioengineering and Bioscience on the Georgia Tech campus.
“One of the unique things about us is, we’re agnostic,” Lawler said. “That is, our startups can come from anywhere. We have companies that have grown out of labs at Georgia State, Alabama State, Emory, and Georgia Tech. And we have interest from entrepreneurs from San Diego, who are considering relocating people from mature biotech markets to our space.”
Ground Floor Companies
Marvin Whiteley wants to help humans win the war against bacteria, and he has a plan, something he’s been cooking up for about 10 years, which has now manifested in his start-up company, SynthBiome, one of the five startups based at BioSpark Labs.
“We can discover a lot of antibiotics in the lab but translating them into the clinic has been a major challenge – antibiotic resistance is the main reason,” said Whiteley, professor in the School of Biological Sciences at Georgia Tech. “Something might work in a test tube easily enough and it might work in a mouse. But the thing is, bacteria know that mice are different - and and so bacteria act differently in mice than in humans.”
SynthBiome was built to help accelerate drug discovery. With that goal in mind, Whiteley and has team set out to develop a better, more effective preclinical model. “We basically learned to let the bacteria tell us what it’s like to be in a human,” Whiteley said. “So, we created a human environment in a test tube.”
Whiteley has said a desire to help people is foundational to his research. He wants to change how successful therapies are made. The same can be said for Dr. Pooja Tiwari, who launched her company, Arnav Biotech, to develop mRNA-based therapeutics and vaccines. Arnav Biotech also serves as a contract researcher and manufacturer, helping other researchers and companies interested in exploring mRNA in their work.
“There are only a handful of people who have deep knowledge of working in mRNA research, and this limits the access to it” said Tiwari, a former postdoctoral researcher at Georgia Tech and Emory. “We’d like to democratize access to mRNA-based therapeutics and vaccines by developing accessible and cost-effective mRNA therapeutics for global needs”.
Arnav – which has RNA right there in the name – in Sanskrit means ‘ocean.’ An ocean has no discernible borders, and Tiwari is working to build a biotech company that eliminates borders in equitable access to mRNA-based therapeutics and vaccines.
With this mission in mind, Arnav is developing mRNA-based, broad-spectrum antivirals as well as vaccines against pandemic potential viruses before the next pandemic hits. Arnav has recently entered in a collaboration with Sartorius BIA Separations, a company based on Slovenia, to advance their mRNA pipeline. While building its own mRNA therapeutics pipeline, Arnav is also helping other scientists explore mRNA as an alternative therapeutic and vaccine platform through its contract services.
“I think of the vaccine scientist who makes his medicine using proteins, but would like to explore the mRNA option,” Tiwari posits. “Maybe he doesn’t want to make the full jump into it. That’s where we come in, helping to drive interest in this field and help that scientist compare his traditional vaccines to see what mRNA vaccines looks like.”
She has all the equipment and instruments that she needs at BioSpark Labs and was one of the first start-ups to put down roots there. So far, it’s been the perfect partnership, Tiwari said, adding, “It kind of feels like BioSpark and Arnav are growing up together.”
News Contact
Writer: Jerry Grillo
Feb. 23, 2023
This news release first appeared in the Chinese Academy of Sciences newsroom, and has been tailored for Georgia Tech readers.
Mycorrhizal symbiosis — a symbiotic relationship that can exist between fungi and plant roots — helps plants expand their root surface area, giving plants greater access to nutrients and water. Although the first and foremost role of mycorrhizal symbiosis is to facilitate plant nutrition, scientists have not been clear how mycorrhizal types mediate the nutrient acquisition and interactions of coexisting trees in forests.
To investigate this crucial relationship, Lingli Liu, a professor at the Institute of Botany of the Chinese Academy of Sciences (IBCAS) led an international, collaborative team, which included School of Biological Sciencesprofessor Lin Jiang. The team studied nutrient acquisition strategies of arbuscular mycorrhizae (AM) and ectomycorrhizal (EcM) trees in the Biodiversity–Ecosystem Functioning (BEF) experiment in a subtropical forest in China, where trees of the two mycorrhizal types were initially evenly planted in mixtures of two, four, eight, or 16 tree species.
The researchers found that as the diversity of species increased, the net primary production (NPP) of EcM trees rapidly decreased, but the NPP of AM trees progressively increased, leading to the sheer dominance (>90%) of AM trees in the highest diversity treatment.
The team's analyses further revealed that differences in mycorrhizal nutrient-acquisition strategies, both nutrient acquisition from soil and nutrient resorption within the plant, contribute to the competitive edge of AM trees over EcM ones.
In addition, analysis of soil microbial communities showed that EcM-tree monocultures have a high abundance of symbiotic fungi, whereas AM-tree monocultures were dominated by saprotrophic and pathogenic fungi.
According to the researchers, as tree richness increased, shifts in microbial communities, particularly a decrease in the relative abundance of Agaricomycetes (mainly EcM fungi), corresponded with a decrease in the NPP of EcM subcommunities, but had a relatively small impact on the NPP of AM subcommunities.
These findings suggest that more efficient nutrient-acquisition strategies, rather than microbial-mediated negative plant-soil feedback, drive the dominance of AM trees in high-diversity ecosystems.
This study, based on the world’s largest forest BEF experiment, provides novel data and an alternative mechanism for explaining why and how AM trees usually dominate in high-diversity subtropical forests.
These findings also have practical implications for species selection in tropical and subtropical reforestation—suggesting it is preferable to plant mixed AM trees, as they have a more efficient nutrient-acquisition strategy than EcM trees.
This study was published as an online cover article in Sciences Advances on Jan. 19 and was funded by the Strategic Priority Research Program of CAS and the National Natural Science Foundation of China.
News Contact
Georgia Tech Editor: Audra Davidson
Communications Officer II
College of Sciences
Feb. 06, 2023
Plants, like animals and people, seek refuge from climate change. And when they move, they take entire ecosystems with them. To understand why and how plants have trekked across landscapes throughout time, researchers at the forefront of conservation are calling for a new framework. The key to protecting biodiversity in the future may be through understanding the past.
Jenny McGuire, assistant professor in the Schools of Biological Sciences and Earth and Atmospheric Sciences at Georgia Tech, spearheaded a special feature on the topic of biodiversity in The Proceedings of the National Academy of Sciences along with colleagues in Texas, Norway, and Argentina. In the special feature, “The Past as a Lens for Biodiversity Conservation on a Dynamically Changing Planet,” McGuire and her collaborators highlight the outstanding questions that must be addressed for successful future conservation efforts. The feature brings together conservation research that illuminates the complex and constantly evolving dynamics brought on by climate change and the ever-shifting ways humans use land. These factors, McGuire said, interact over time to create dynamic changes and illustrate the need to incorporate temporal perspectives into conservation strategies by looking deep into the past.
One example of this work highlighted in the journal is McGuire’s research about plants in North America, which investigates how and why they’ve moved across geography over time, where they’re heading, and why it’s important.
“Plants are shifting their geographic ranges, and this is happening whether we realize it or not,” McGuire said. “As seeds fall or are transported to distant places, the likelihood that the plant’s seed is going to be able to survive and grow is changing as climates are changing. Studying plants’ niche dynamics over thousands of years can help us understand how species adapt to climate change and can teach us how to protect and maintain biodiversity in the face of rapid climate change to come.”
Climate Fidelity: A New Metric for Understanding Vulnerability
The first step is to understand which type of plants exhibit what McGuire terms “climate fidelity,” and which do not. If a plant has climate fidelity, it means that the plant stays loyal to its preferred climatic niche, often migrating across geographies over thousands of years to keep up with its ideal habitat. Plants that don’t exhibit climate fidelity tend to adapt locally in the face of climate change. Being loyal to one’s climate, it turns out, doesn’t necessarily mean being loyal to a particular place.
To investigate the case of trees, McGuire and former Georgia Tech postdoctoral scholar Yue Wang (associate professor in the School of Ecology at Sun Yat-sen University in China) studied pollen data from the Neotoma Paleoecology Database, which contains pollen fossil data from sediment cores across North America. Each sediment core is sampled, layer by layer, producing a series of pollen data from different times throughout history. The data also contains breakdowns of the relative abundance of different types of plants represented by the pollen types – pine versus oak versus grass, for example – painting a picture of what types of plants were present in that location and when.
McGuire and Wang looked at data from 13,240 fossil pollen samples taken from 337 locations across the entirety of North America. For each of the 16 major plant taxa in North America, they divided the pollen data into six distinct chunks or “bins” of time of 4,000 years, starting from 18,000 years ago up to the present day. Wang used the data to identify all climate sites containing fossil pollen for any individual type of tree – such as oak, for example – for each period. Then, Wang looked at how each tree’s climate changed from one period to the next. Wang did this by comparing the locations of pollen types between adjacent time periods, which enabled the team to identify how and why each type of tree’s climate changed over time.
“This process allowed us to see the climate fidelity of these different plant taxa, showing that certain plants maintain very consistent climatic niches, even when climate is changing rapidly,” Wang said.
For example, their findings showed that when North American glaciers were retreating 18,000 years ago, spruce and alder trees moved northward to maintain the cool temperatures of their habitats.
Crucially, McGuire and Wang found that most plant species in North America have exhibited long-term climate fidelity over the past 18,000 years. They also found that plants that migrated farther did a better job of tracking climate during periods of change.
But some plants fared better than others. For example, the small seeds of willow trees can fly over long distances – enabling them to track their preferred climates very effectively. But the large seeds of ash trees, for example, can only be dispersed short distances from parent trees, hindering their ability to track climate. Habitat disruptions from humans could make it even more difficult for ash trees to be able to take hold in new regions. If there are no adjacent habitats for ash trees, their seeds are under pressure to move even farther – a particular challenge for ash, which slows their migration movements even more.
Protecting the Fabric of Life
On the bright side, by identifying which plants have historically been most sensitive to changing climates, McGuire and Wang’s research can help conservation organizations like The Nature Conservancy prioritize land where biodiversity is most vulnerable to climate change.
As a final step, McGuire and Wang identified “climate fidelity hotspots,” regions that have historically exhibited strong climate fidelity whose plants will most urgently need to move as their climates change. They compared these hotspots to climate-resilient regions identified by The Nature Conservancy that could serve as refuge areas for those plants. While plants in these resilient regions can initially adapt to impending climate change by shifting their distributions locally, the plants will likely face major challenges when a region’s climate change capacity is exceeded due to lack of connectivity and habitat disruptions from humans. Refining these priorities helps stakeholders identify efficient strategies for allowing the fabric of life to thrive.
“I think that understanding climate fidelity, while a new and different idea, will be very important going forward, especially when thinking about how to prioritize protecting different plants in the face of climate change,” McGuire said. “It is important to be able to see that some plants and animals are more vulnerable to climate change, and this information can help build stronger strategies for protecting the biodiversity on the planet.”
Citation: Yue Wang, Silvia Pineda-Munoz, and Jenny L. McGuire, "Plants maintain climate fidelity in the face of dynamic climate change." PNAS (2023).
DOI: doi.org/10.1073/pnas.2201946119
News Contact
Catherine Barzler, Senior Research Writer/Editor
Pagination
- Previous page
- Page 22
- Next page