May. 09, 2024
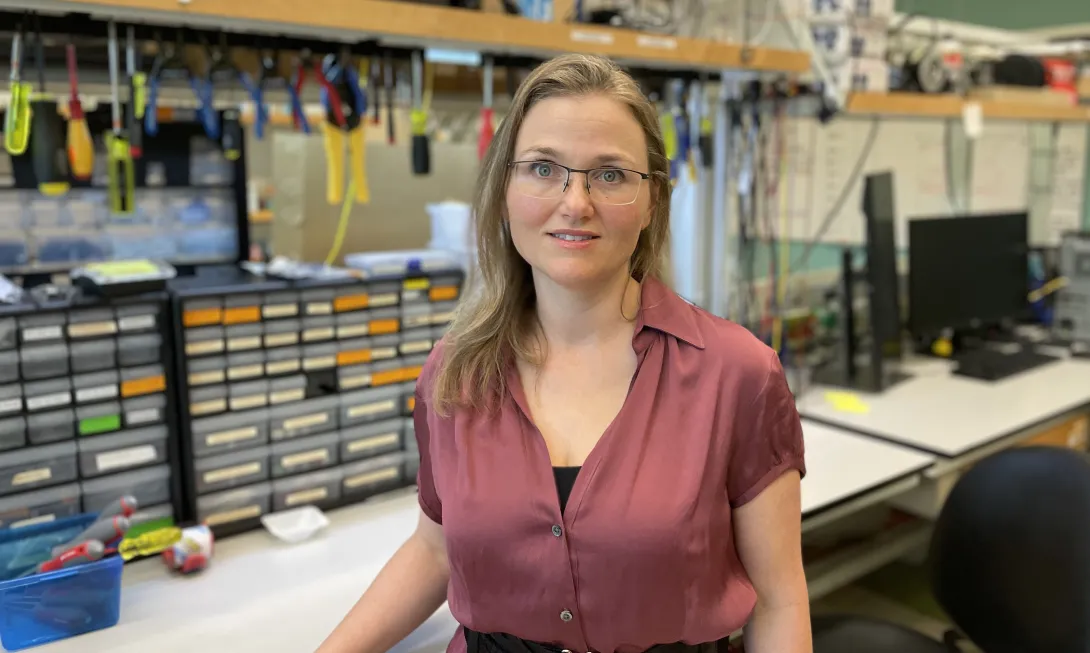
Biomedical engineer Annabelle Singer has spent the past decade developing a noninvasive therapy for Alzheimer’s disease that uses flickering lights and rhythmic tones to modulate brain waves. Now she has discovered that the technique, known as flicker, also could benefit patients with a host of other neurological disorders, from epilepsy to multiple sclerosis.
Previously, Singer and her collaborators demonstrated that the lights and sounds, delivered to patients through goggles and headphones, have beneficial effects. Flicker has been successful in animal studies and in early human feasibility trials, where it was tested for safety, tolerance, and patient adherence.
Now, thanks to a clinical trial for people with epilepsy, the researchers quantified flicker’s effects with unprecedented precision. They also made an unexpected, but encouraging, discovery: The treatment reduced interictal epileptiform discharges (IEDs) in the brain.
These large, intermittent electrophysiological events are observed between seizures in people with epilepsy. They appear as sharp spikes on an EEG readout.
“What’s interesting about these IEDs is that they don’t just occur in epilepsy,” said Singer, McCamish Foundation Early Career Professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. “They occur in autism, multiple sclerosis, Alzheimer’s, and other neurological disorders, too.” And IEDs disrupt normal brain function, causing memory impairment.
Singer and her team published their findings recently in Nature Communications.
The Rhythm in Our Heads
Inside the brain are elaborate symphonies of electrical activity: brain waves, or oscillations, that compose our memories, thoughts, and emotions. Singer wants to modulate those oscillations for therapeutic purposes.
At specific frequencies of light and sound, the flicker treatment can induce gamma oscillations in mice. This helps the brain recruit microglia, cells responsible for removing beta amyloid, which is believed to play a central role in Alzheimer’s pathology. Part of the work is in recording what’s happening in the brain during treatment to verify how it’s working.
The patients in the trial were under the care of physician Jon Willie at the Emory University Hospital Epilepsy Monitoring Unit. (Willie, co-corresponding author of the study with Singer, is now at Washington University in St. Louis.) They were awaiting surgery to remove an area of the brain where seizures occur. Before that could happen, they had to undergo intracranial seizure monitoring — recording electrodes are placed in the brain to pinpoint the seizure onset zone and determine exactly which tissue should be removed. Then, patients and their care team wait for a seizure to happen. It can take days.
“In human studies, we’ve used noninvasive methods like functional MRI or scalp EEG, but they have real downsides in terms of resolution,” Singer said. “Working with these patients was a game changer. These are people with treatment-resistant epilepsy, which means that drugs aren’t working for them.”
Pathway to Healing
Singer’s team recruited 19 patients. Lead author of the study, Lou Blanpain, a former Ph.D. student in Singer’s lab and now a medical student at Emory, went from patient to patient with the flicker stimulation and recording equipment.
“Because these patients already had recording probes implanted for clinical reasons, we were able to record directly from the brain,” Singer said. “We’ve never been able to get recordings of this quality during flicker treatment before.”
As the researchers expected, flicker modulated the visual and auditory brain regions that respond strongly to stimuli. But it also reached deeper, into the medial temporal lobe and prefrontal cortex, brain regions crucial for memory. And across the brain, in regions Singer hadn’t fully explored before, she found IEDs were decreasing.
“That has important implications for whether flicker is therapeutically relevant for people with Alzheimer’s, but also in general if we want to target anything beyond the primary sensory regions,” she said. “All of this points to the potential use of flicker in a lot of different contexts. Going forward, we’re definitely going to look at other conditions and other potential implications.”
Citation: Lou T. Blanpain, Eric R. Cole, Emily Chen, James K. Park, Michael Y. Walelign, Robert E. Gross, Brian T. Cabaniss, Jon T. Willie, Annabelle C. Singer. “Multisensory Flicker Modulates Widespread Brain Networks and Reduces Interictal Epileptiform Discharges,” Nature Communications.
Funding: National Institutes of Health (R01 NS109226, RF1NS109226, RF1AG078736, R01 MH120194, P41 EB018783, MH12019), DARPA, McCamish Foundation, Packard Foundation.
Competing interests: Annabelle Singer owns shares in Cognito Therapeutics, which aims to develop gamma stimulation-related products. These conflicts are managed by Georgia Tech’s Office of Research Integrity Assurance.
News Contact
Jerry Grillo
May. 06, 2024
Cardiologists and surgeons could soon have a new mobile augmented reality (AR) tool to improve collaboration in surgical planning.
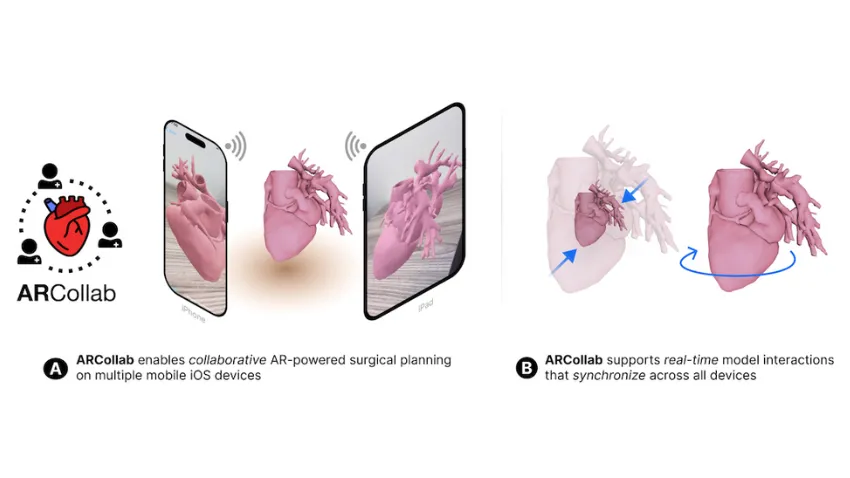
ARCollab is an iOS AR application designed for doctors to interact with patient-specific 3D heart models in a shared environment. It is the first surgical planning tool that uses multi-user mobile AR in iOS.
The application’s collaborative feature overcomes limitations in traditional surgical modeling and planning methods. This offers patients better, personalized care from doctors who plan and collaborate with the tool.
Georgia Tech researchers partnered with Children’s Healthcare of Atlanta (CHOA) in ARCollab’s development. Pratham Mehta, a computer science major, led the group’s research.
“We have conducted two trips to CHOA for usability evaluations with cardiologists and surgeons. The overall feedback from ARCollab users has been positive,” Mehta said.
“They all enjoyed experimenting with it and collaborating with other users. They also felt like it had the potential to be useful in surgical planning.”
ARCollab’s collaborative environment is the tool’s most novel feature. It allows surgical teams to study and plan together in a virtual workspace, regardless of location.
ARCollab supports a toolbox of features for doctors to inspect and interact with their patients' AR heart models. With a few finger gestures, users can scale and rotate, “slice” into the model, and modify a slicing plane to view omnidirectional cross-sections of the heart.
Developing ARCollab on iOS works twofold. This streamlines deployment and accessibility by making it available on the iOS App Store and Apple devices. Building ARCollab on Apple’s peer-to-peer network framework ensures the functionality of the AR components. It also lessens the learning curve, especially for experienced AR users.
ARCollab overcomes traditional surgical planning practices of using physical heart models. Producing physical models is time-consuming, resource-intensive, and irreversible compared to digital models. It is also difficult for surgical teams to plan together since they are limited to studying a single physical model.
Digital and AR modeling is growing as an alternative to physical models. CardiacAR is one such tool the group has already created.
However, digital platforms lack multi-user features essential for surgical teams to collaborate during planning. ARCollab’s multi-user workspace progresses the technology’s potential as a mass replacement for physical modeling.
“Over the past year and a half, we have been working on incorporating collaboration into our prior work with CardiacAR,” Mehta said.
“This involved completely changing the codebase, rebuilding the entire app and its features from the ground up in a newer AR framework that was better suited for collaboration and future development.”
Its interactive and visualization features, along with its novelty and innovation, led the Conference on Human Factors in Computing Systems (CHI 2024) to accept ARCollab for presentation. The conference occurs May 11-16 in Honolulu.
CHI is considered the most prestigious conference for human-computer interaction and one of the top-ranked conferences in computer science.
M.S. student Harsha Karanth and alumnus Alex Yang (CS 2022, M.S. CS 2023) co-authored the paper with Mehta. They study under Polo Chau, an associate professor in the School of Computational Science and Engineering.
The Georgia Tech group partnered with Timothy Slesnick and Fawwaz Shaw from CHOA on ARCollab’s development.
“Working with the doctors and having them test out versions of our application and give us feedback has been the most important part of the collaboration with CHOA,” Mehta said.
“These medical professionals are experts in their field. We want to make sure to have features that they want and need, and that would make their job easier.”
News Contact
Bryant Wine, Communications Officer
bryant.wine@cc.gatech.edu
May. 02, 2024
The University System of Georgia Board of Regents has approved a new Neuroscience and Neurotechnology Ph.D. Program at Georgia Tech.
The interdisciplinary degree is a joint effort across the Colleges of Sciences, Computing, and Engineering. The program expects to enroll its first graduate students in Fall 2025, pending approval by the Southern Association of Colleges and Schools Commission on Colleges.
The Institute Curriculum Committee has also approved a new Minor in Neuroscience, set to become available in the Georgia Tech 2024-2025 Catalog.
B.S. in Neuroscience
The Ph.D. and Minor offerings build on the recently launched Neuro Next Initiative in Research, and the established Undergraduate Program in Neuroscience, respectively.
Approved by the Board of Regents in 2017, the interdisciplinary B.S. in Neuroscience degree in the College of Sciences enrolled more than 400 undergraduate students in 2022, and has been the fastest growing undergraduate major at Georgia Tech.
The B.S. in Neuroscience is also key to a strong ecosystem of undergraduate neuroscience education across the state, which includes peer programs at Mercer University, Augusta University, Georgia State University, Agnes Scott College, and Emory University.
Ph.D. in Neuroscience and Neurotechnology
The new doctoral degree will provide a path for the rapidly growing pipeline of in-state neuroscience undergraduate students and young alumni — while also welcoming a wider slate of graduate researchers to campus.
The Ph.D. Program’s mission is focused on educating students to advance the field of neuroscience through an interdisciplinary approach, with scientists and engineers of different backgrounds — ultimately integrating neuroscience research and technological development to study all levels of nervous system function.
Biological Sciences Professor Lewis A. Wheaton, who chaired the Ph.D. Program Planning Committee, shares that a cohort model will fuse “experimental and quantitative skill development, creating opportunities for students to work in science and engineering labs to promote collaborations, while also fostering a program and community that’s unique to the state and against national peer offerings.”
Expanding innovation — and impact
Wheaton explains that the new Ph.D. aims to equip graduates for a wide range of employment opportunities and growing specializations, including computational neuroscience, neurorehabilitation, cultural and social neuroscience, neuroimaging, cognitive and behavioral neuroscience, developmental neuroscience, and neurolinguistics.
The new degree will also help meet the country’s growing demand for a neuro-centric workforce. According to the U.S. Bureau of Labor Statistics, job growth for medical scientists (including neuroscientists) tracked around 13% between 2012 and 2022, faster than the average for all tracked occupations.
Wheaton adds that the program will equip neuroscientists to conduct research that can significantly improve lives.
Seeking students
The Planning Committee anticipates a tentative February 1, 2025 application deadline for Fall 2025 enrollments — and encourages students with the following interests to learn more and apply in the coming school year:
- Developing deeper quantitative, computing and/or engineering skills to make scientific discoveries that support innovations in neuroscience
- A clear, comprehensive understanding of the nervous system at all scales from molecular to systems
- Understanding how to use and innovate new tools and approaches to investigate the nervous system at all levels
- Becoming uniquely qualified to translate knowledge across neuroscience and related disciplines to create new knowledge in their professional pursuits
Director search
The participating Colleges will soon conduct a search for a program director, engaging a tenured member of the Georgia Tech faculty to serve as the new program’s administrator. A graduate program committee composed of five faculty members and mentors across the Colleges of Sciences, Computing, and Engineering, will also be created.
During their April 2024 meeting, Regents also announced budget approvals and tuition changes for Georgia's 26 member institutions.
The Ph.D. Program Planning Committee included the following faculty:
- Lewis Wheaton (Committee Chair, Biological Sciences)
- Constantine Dovrolis (Computer Science)
- Christopher Rozell (Electrical and Computer Engineering)
- Eric Schumacher (Psychology)
- Garrett Stanley (Biomedical Engineering)
- David Collard (College of Sciences Office of the Dean)
News Contact
Programs:
- Ph.D. in Neuroscience and Neurotechnology
Contact Professor Lewis Wheaton, Planning Committee Chair - Undergraduate Program in Neuroscience
- Minor in Neuroscience
- Georgia Tech Neuro and Neuro Next
Press Contact:
Jess Hunt-Ralston
Director of Communications
College of Sciences at Georgia Tech
Neuro Next Initiative:
Sarah Peterson
Program Manager
GT Neuro
Audra Davidson
Research Communications Program Manager
Neuro Next Initiative at Georgia Tech
Apr. 29, 2024
From her home more than 800 miles away, Georgia Tech online master's student Jasmine Tata is monitoring fish in aquariums at Georgia Tech.
Tata is a New York-based QA analyst and project manager. She started the Online Master of Science in Computer Science (OMSCS) program in Fall 2022 and joined FishStalkers last year.
The student-led research program is part of the School of Biological Sciences' McGrath Lab. Its researchers use machine learning, computer vision, and other technologies to better understand the evolution of animal behaviors.
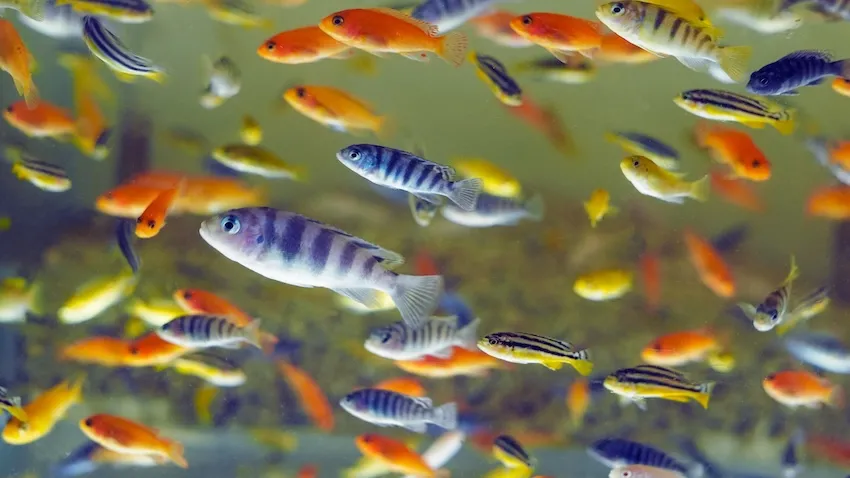
One of the lab's research projects studies Lake Malawi cichlids to explore connections between observed behavior and brain function.
The FishStalkers are vital to the project. They collect video, depth, and other data from individual fish using Raspberry Pi single-board computers. This information, coupled with open-source code they developed, allows the group to track, monitor, and classify the behaviors of a fish as it builds and maintains its bower, which is a sand structure these cichlids use to attract mates.
Along with monitoring the research tanks, Tata's contributions include improving the automated collection and analysis of data streaming from the Pis. She's also helping to adapt the data pipeline to work with yellow-head, orange-cap, and other cichlid species.
[RELATED: Georgia Tech's OMSCS Program Celebrates 10th Anniversary]
"I've enjoyed learning more about new problems in a relatively unfamiliar field. In a pure computer science-focused lab, I never would experience the frustrations of data collection that come with biological subjects," said Tata.
"The fish builds bowers on its own schedule, and data collection must accurately capture this, regardless of weekends or holidays."
Tata says her experience with FishStalkers has given her new ideas about presenting data to non-technical team members. The team uses a spreadsheet integrated with data collection scripts running on the Raspberry Pis. The spreadsheet allows someone without technical knowledge to pause, upload data, or start new trials simply by toggling a dropdown.
"This has given me a lot of ideas about how to meet people where they are in terms of technical skills when it comes to user interface design and has encouraged me to learn more about human-computer interaction," said Tata.
Tata learned about the FishStalkers research group when its founder, Breanna Shi, reached out through the OMSCS Slack study channel. Shi developed the group through Georgia Tech's Vertically Integrated Projects (VIP) program as a mentorship program.
"Given their real-world computer science experience, I wanted to see if there were OMSCS students interested in collaborating on FishStalkers projects and assisting in the mentorship of undergraduate researchers," said Shi.
Shi is a third-year Ph.D. student studying bioinformatics with minors in machine learning and higher education. She created FishStalkers as a mentorship program because she recognized that undergraduate and masters-level students could feel less valued or isolated in research environments.
"The FishStalkers model empowers all its researchers with the respect and responsibility as a full team member. Whether it's your first week as a FishStalker or your last, you will complete tasks that benefit the research team and yourself," said Shi.
[RELATED: Women-Centered Mentorship Provides Empowerment to Conquer Ph.D.]
Tata's experience in the business world made her a good fit for the FishStalkers program. Shi says Tata contributes valuable insight to the group as a mentor because most students approach the program from a purely academic viewpoint.
"Jasmine, like other OMSCS students, works full-time and attends the OMSCS program part-time. Her roles as a project manager and a software QA analyst allow her to contribute a unique perspective to the FishStalkers group," said Shi.
In addition to sharing her experience mentoring two OMSCS students this semester, Tata has helped Shi overcome some of the inherent challenges of long-distance collaboration. These include creating a sense of interpersonal connection among in-person and remote research team members.
Group meetings host a virtual link to enhance the online research experience. Every member provides progress updates during the sessions. The researchers also virtually check in and out of their research hours in a shared group chat and describe the work completed during their check-out.
"FishStalkers also runs a monthly lab-buddy program where a researcher is paired with a new buddy each month to schedule a 30-minute meeting to chat and learn about each other's work," said Shi.
"These strategies benefit OMSCS students in our group and provide a positive research environment for junior researchers. We seek to incorporate innovative strategies to create an accessible research environment for all students interested in participating in our research," said Shi.
FishStalkers has been such a success that Shi is expanding the model. This fall, Shi will work with OMSCS Executive Director David Joyner and OMSCS Associate Director of Research Nick Lytle to connect OMSCS students with interdisciplinary research projects in labs across campus.
"My role will be to establish relationships between data collectors and data analyzers to provide a service to non-technical labs across campus and a valuable research experience for OMSCS students," said Shi.
"We will be building from my existing work in image processing in the McGrath Lab and expanding to other labs with data analysis needs. I am very excited to have the experience of growing as a collaborator."
News Contact
Ben Snedeker, Communications Manager
Georgia Tech College of Computing
Apr. 23, 2024
This article was first published in the University of Illinois Urbana-Champaign newsroom. Read the full story here.
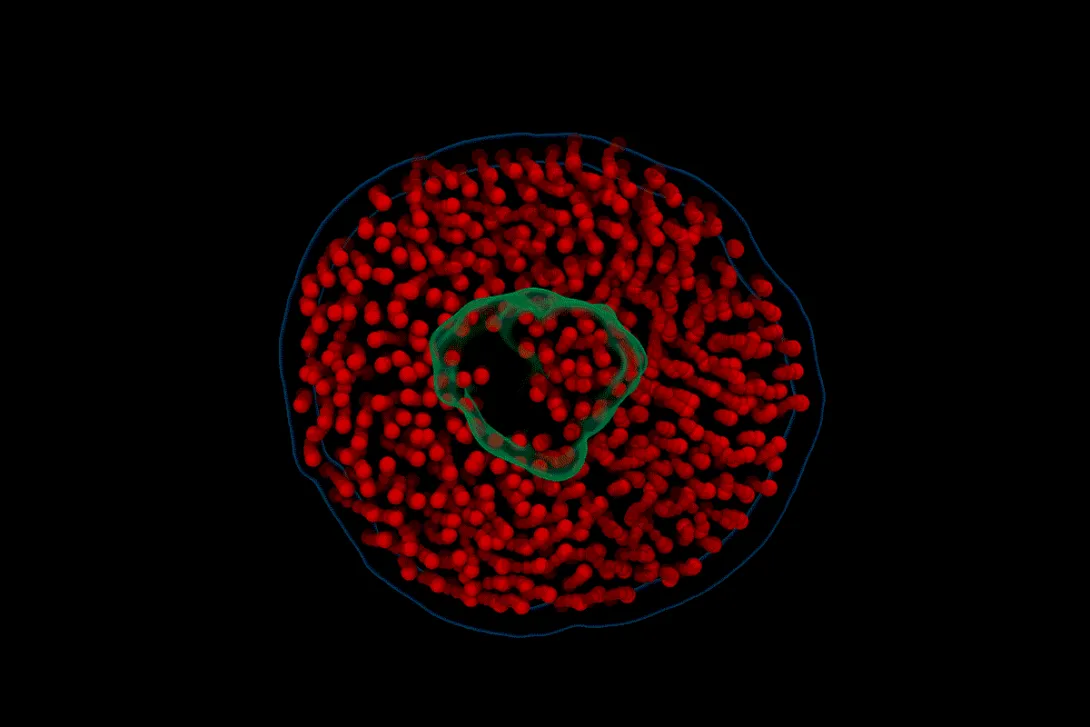
Researchers at Georgia Institute of Technology and the University of Illinois Urbana-Champaign have developed a first-of-its-kind technique called electron videography to capture moving images at the molecular scale. In the first demonstration of the technique, the team took a microscopic moving picture of the delicate dance between proteins and lipids found in cell membranes. The study, “Electron videography of a lipid–protein tango” was published last week in the journal Science Advances.
"This is the first time we are looking at a protein on an individual scale and haven't frozen it or tagged it," says Aditi Das, a corresponding author and associate professor in the School of Chemistry and Biochemistry at Georgia Tech.
Electron microscopy techniques image at the molecular or atomic scale, yielding detailed, nanometer-scale pictures. However, they often rely on samples that have been frozen or fixed in place, leaving scientists to try to infer how molecules move and interact — like trying to map the choreography of a dance sequence from a single frame of film.
"Usually, we have to crystalize or freeze a protein, which poses challenges in capturing high-resolution images of flexible proteins. Alternately, some techniques use a molecular tag that we track, rather than watching the protein itself,” Das says. “In this study we are seeing the protein as it is, behaving how it does in a liquid environment, and seeing how lipids and proteins interact with each other."
The technique can be used to study the dynamics of other biomolecules, breaking free of constraints that have limited microscopy to still images of fixed molecules. In this study, the team examined nanoscale discs of lipid membranes and how they interacted with proteins normally found on the surface of or embedded in cell membranes.
These membrane proteins are significant for medical treatments, and are involved in processes including muscle contraction, brain function, and immune system functions. Moving forward, the researchers plan to use their electron videography technique to study other types of membrane proteins and other classes of molecules and nanomaterials.
News Contact
Contact:
Jess-Hunt Ralston
Director of Communications
College of Sciences
Georgia Tech
Apr. 18, 2024
Aaron Levine, associate dean for research and outreach in the Ivan Allen College of Liberal Arts, has been named a fellow of the American Association for the Advancement of Science (AAAS), the world’s largest multidisciplinary scientific society.
Levine is one of 502 people named to the AAAS Fellows Class of 2023, an honor the society has been awarding scientists, engineers, and innovators since 1874 for achievements and efforts on behalf of the advancement of science and its applications. AAAS Fellows are recognized for outstanding contributions to research, teaching, technology, and science communication.
“I am deeply honored to receive this recognition from an organization that has supported and inspired me since I joined as a graduate student in 2006,” said Levine. “I am also encouraged by this acknowledgement that policy and ethics play a key role in bringing groundbreaking biomedical technologies to the people who need them.”
The organization chose Levine, who is also a professor in the School of Public Policy, for his contributions to biomedical research policy — including advancing understanding of how policy debates influence contentious areas of research. His work is at the intersection of ethics, policy, and biomedical research.
“I first became interested in bioethics and science policy while working on the human genome project and witnessing the ethical and policy issues that arose,” said Levine. “I believe addressing societal issues associated with emerging biomedical technologies is critical for these advances to reach their full potential.”
Levine’s work focuses on the development and oversight of biomedical research and health care areas such as stem cell treatments, assisted reproductive technology, fetal tissue research, and CRISPR.
The author of Cloning: A Beginner’s Guide, an accessible introduction to the science of cloning and the ethical and policy controversies this science inspires, Levine also has a longstanding interest in science communication. He was a member of the 2019-20 cohort of AAAS’ Alan I. Leshner Leadership Institute Public Engagement Fellows.
In addition to his duties in the School and College, Levine also leads ethics and policy research for the National Science Foundation Engineering Research Center for Cell Manufacturing Technologies (CMaT). From 2017 to 2022, he served as CMaT’s co-director for engineering workforce development, helping guide efforts to produce a diverse, well-trained workforce for the biomanufacturing industry.
Levine holds a Ph.D. in public affairs from Princeton University and a master of philosophy from the University of Cambridge, where he was a Churchill Scholar. He earned a bachelor of science in biology from the University of North Carolina at Chapel Hill, where he was a Morehead Scholar.
News Contact
Stephanie N. Kadel
Ivan Allen College of Liberal Arts
Mar. 18, 2024
Computer science educators will soon gain valuable insights from computational epidemiology courses, like one offered at Georgia Tech.
B. Aditya Prakash is part of a research group that will host a workshop on how topics from computational epidemiology can enhance computer science classes.
These lessons would produce computer science graduates with improved skills in data science, modeling, simulation, artificial intelligence (AI), and machine learning (ML).
Because epidemics transcend the sphere of public health, these topics would groom computer scientists versed in issues from social, financial, and political domains.
The group’s virtual workshop takes place on March 20 at the technical symposium for the Special Interest Group on Computer Science Education (SIGCSE). SIGCSE is one of 38 special interest groups of the Association for Computing Machinery (ACM). ACM is the world’s largest scientific and educational computing society.
“We decided to do a tutorial at SIGCSE because we believe that computational epidemiology concepts would be very useful in general computer science courses,” said Prakash, an associate professor in the School of Computational Science and Engineering (CSE).
“We want to give an introduction to concepts, like what computational epidemiology is, and how topics, such as algorithms and simulations, can be integrated into computer science courses.”
Prakash kicks off the workshop with an overview of computational epidemiology. He will use examples from his CSE 8803: Data Science for Epidemiology course to introduce basic concepts.
This overview includes a survey of models used to describe behavior of diseases. Models serve as foundations that run simulations, ultimately testing hypotheses and making predictions regarding disease spread and impact.
Prakash will explain the different kinds of models used in epidemiology, such as traditional mechanistic models and more recent ML and AI based models.
Prakash’s discussion includes modeling used in recent epidemics like Covid-19, Zika, H1N1 bird flu, and Ebola. He will also cover examples from the 19th and 20th centuries to illustrate how epidemiology has advanced using data science and computation.
“I strongly believe that data and computation have a very important role to play in the future of epidemiology and public health is computational,” Prakash said.
“My course and these workshops give that viewpoint, and provide a broad framework of data science and computational thinking that can be useful.”
While humankind has studied disease transmission for millennia, computational epidemiology is a new approach to understanding how diseases can spread throughout communities.
The Covid-19 pandemic helped bring computational epidemiology to the forefront of public awareness. This exposure has led to greater demand for further application from computer science education.
Prakash joins Baltazar Espinoza and Natarajan Meghanathan in the workshop presentation. Espinoza is a research assistant professor at the University of Virginia. Meghanathan is a professor at Jackson State University.
The group is connected through Global Pervasive Computational Epidemiology (GPCE). GPCE is a partnership of 13 institutions aimed at advancing computational foundations, engineering principles, and technologies of computational epidemiology.
The National Science Foundation (NSF) supports GPCE through the Expeditions in Computing program. Prakash himself is principal investigator of other NSF-funded grants in which material from these projects appear in his workshop presentation.
[Related: Researchers to Lead Paradigm Shift in Pandemic Prevention with NSF Grant]
Outreach and broadening participation in computing are tenets of Prakash and GPCE because of how widely epidemics can reach. The SIGCSE workshop is one way that the group employs educational programs to train the next generation of scientists around the globe.
“Algorithms, machine learning, and other topics are fundamental graduate and undergraduate computer science courses nowadays,” Prakash said.
“Using examples like projects, homework questions, and data sets, we want to show that the topics and ideas from computational epidemiology help students see a future where they apply their computer science education to pressing, real world challenges.”
News Contact
Bryant Wine, Communications Officer
bryant.wine@cc.gatech.edu
Mar. 18, 2024
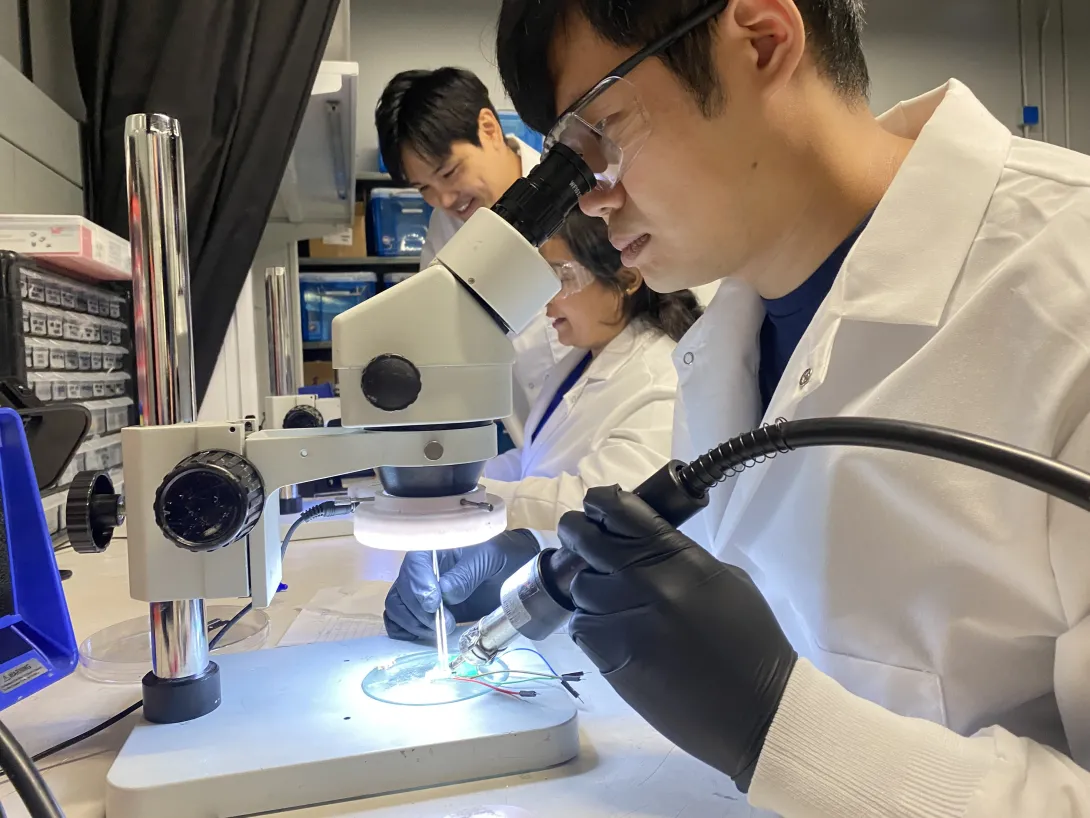
Researchers with Georgia Tech and Emory University are field testing a new device that could help protect people who work outside from heat related injury. It’s a skin patch you can wear while working that sends detailed information to a smartphone or other device about important health markers like skin hydration and body temperature. The device takes different measurements than health wearables on the market currently and will be paired with an artificial intelligence program to predict health hazards. The team is calling the device BioPatch, and it’s being put to the test with landscaping crews. Researchers hope use of the device can guide better decisions about working in the heat.
The project involves collaboration between principal investigators Vicki Hertzberg from Emory University, W. Hong Yeo from Georgia Tech, and Li Xiong from Emory University. Their expertise spans statistics, mechanical and biomedical engineering, and computer science, respectively. Roxana Chicas of the Emory School of Nursing and Jeff Sands of the Emory School of Medicine, along with members of the Farmworker Association of Florida, are also part of the team. This video shows the device and data collection during a key component of testing during the summer.
Mar. 12, 2024
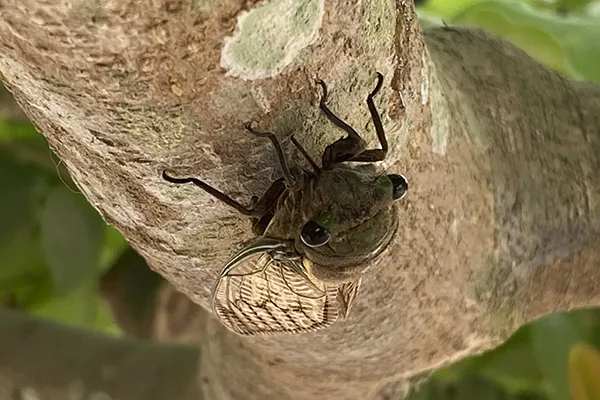
Cicadas are the soundtrack of summer, but their pee is more special than their music. Rather than sprinkling droplets, they emit jets of urine from their small frames. For years, Georgia Tech researchers have wanted to understand the cicada’s unique urination.
Cicadas are the soundtrack of summer, but their pee is more special than their music. Rather than sprinkling droplets, they emit jets of urine from their small frames. For years, Georgia Tech researchers have wanted to understand the cicada’s unique urination.
Saad Bhamla, an assistant professor in the School of Chemical and Biochemical Engineering, and his research group hoped for an opportunity to study a cicada’s fluid excretion. However, while cicadas are easily heard, they hide in trees, making them hard to observe. As such, seeing a cicada pee is an event. Bhamla’s team had only watched the process on YouTube.
Then, while doing fieldwork in Peru, the team got lucky: They saw numerous cicadas in a tree, peeing.
Read more about what they discovered at Georgia Tech Research News.
News Contact
Tess Malone, Senior Research Writer/Editor
tess.malone@gatech.edu
Jan. 23, 2024
Valeria Tohver Milam leads the Macromolecular Materials at Biotic and Abiotic Interfaces research initiative for the Institute for Materials (IMat) and the Parker H. Petit Institute for Bioengineering and Biosciences at Georgia Tech. In this role, she is working to build an inclusive and active community across and beyond Georgia Tech to identify emerging research directions in macromolecular materials for biological and nonbiological applications. Milam is an associate professor in Materials Science and Engineering and a program faculty member of the Bioengineering graduate program at Georgia Tech.
In this brief Q&A, Milam discusses her research focus, how it relates to materials research, and the impact of this initiative.
What is your field of expertise and at what point in your life did you first become interested in this area?
My field of expertise lies in bio-inspired materials science and engineering. Natural macromolecular components of biological systems such as cell receptors or antibodies rely on recognition-based binding events to, for example, allow a cell to take up particular nutrients or to neutralize a specific pathogen threat. Inspired by nature’s capabilities, my group’s research strives to identify and study synthetic macromolecular materials with bio-inspired compositions and self-folded structures. I first became interested in using DNA for its recognition capabilities during my postdoc at the University of Pennsylvania. For the first several years as an assistant professor at Georgia Tech, my group used DNA duplexes as a temporary glue between particle surfaces. Our more recent efforts focus on finding oligonucleotides to function as ligands or capture agents for a specific biological or nonbiological target.
What questions or challenges sparked your current materials research?
Polymers or macromolecules hold a lot of promise as a class of materials for various applications. Synthetic macromolecules, however, pose a lot of synthesis and post-use challenges that can hinder the discovery and practical use of novel macromolecular chemistries. Natural polymers such as oligonucleotides and proteins, on the other hand, have their own elegant synthesis and degradation pathways. To promote discovery of novel macromolecular materials, my group uses nature’s reagents and building blocks to synthesize numerous artificial biopolymer candidates. Since we do not start with any sequence design rules, we rely on maximizing the composition diversity of these artificial biopolymers. We then test all candidates collectively to efficiently choose ones with the desired functionality.
Why is your initiative important to the development of Georgia Tech’s Materials research strategy?
One of the challenges to discovering macromolecular systems that are both novel and practical is the lack of design rules. For example, how does one choose the right number and composition of repeat units for a macromolecule that binds to a particular material surface or to a particular biological target. If you can take advantage of nature’s building blocks and enzymes, then you can explore a wide chemical combinatorial space without having to follow any prerequisite design rules. Better yet, you can then use your initial findings to come up with design rules to explore additional, possibly better macromolecular candidates. This approach to macromolecule discovery is inherently interdisciplinary since one must combine or adapt techniques and approaches developed by biologists, polymer scientists, and materials engineers. Thus, Georgia Tech is a great place to foster this interdisciplinary strategy to research.
What are the broader global and social benefits of the research you and your team conduct?
In addition to training members of our future workforce with interdisciplinary skill sets, we want to carve out a pathway to designing, synthesizing and using environmentally friendly, multiuse macromolecules with commercial promise.
What are your plans for engaging a wider GT faculty pool with IMat research?
Currently, we are primarily in the brainstorming stage. To this end, I am engaging with science and engineering faculty at GT as well as Emory. As cross-disciplinary ideas start to brew, we will work towards multi-PI funding opportunities that engage the broader GT faculty and community.
News Contact
Amelia Neumeister
Research Communications
Pagination
- Previous page
- Page 20
- Next page