Jul. 15, 2024
Hepatic, or liver, disease affects more than 100 million people in the U.S. About 4.5 million adults (1.8%) have been diagnosed with liver disease, but it is estimated that between 80 and 100 million adults in the U.S. have undiagnosed fatty liver disease in varying stages. Over time, undiagnosed and untreated hepatic diseases can lead to cirrhosis, a severe scarring of the liver that cannot be reversed.
Most hepatic diseases are chronic conditions that will be present over the life of the patient, but early detection improves overall health and the ability to manage specific conditions over time. Additionally, assessing patients over time allows for effective treatments to be adjusted as necessary. The standard protocol for diagnosis, as well as follow-up tissue assessment, is a biopsy after the return of an abnormal blood test, but biopsies are time-consuming and pose risks for the patient. Several non-invasive imaging techniques have been developed to assess the stiffness of liver tissue, an indication of scarring, including magnetic resonance elastography (MRE).
MRE combines elements of ultrasound and MRI imaging to create a visual map showing gradients of stiffness throughout the liver and is increasingly used to diagnose hepatic issues. MRE exams, however, can fail for many reasons, including patient motion, patient physiology, imaging issues, and mechanical issues such as improper wave generation or propagation in the liver. Determining the success of MRE exams depends on visual inspection of technologists and radiologists. With increasing work demands and workforce shortages, providing an accurate, automated way to classify image quality will create a streamlined approach and reduce the need for repeat scans.
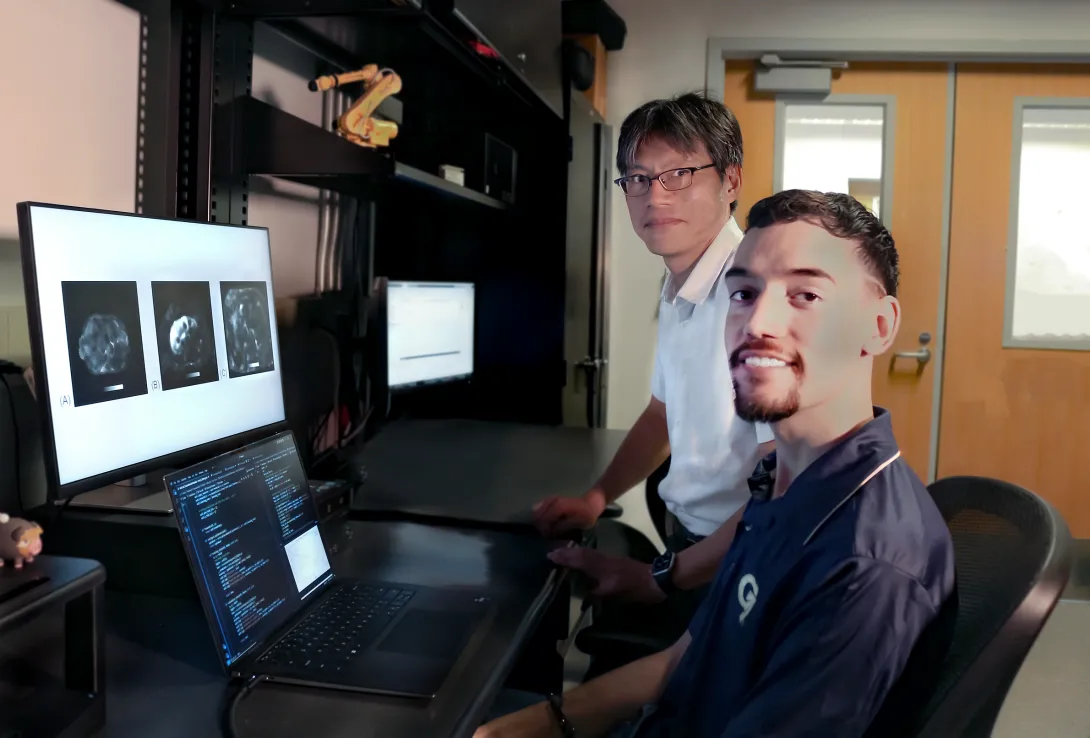
Professor Jun Ueda in the George W. Woodruff School of Mechanical Engineering and robotics Ph.D. student Heriberto Nieves, working with a team from the Icahn School of Medicine at Mount Sinai, have successfully applied deep learning techniques for accurate, automated quality control image assessment. The research, “Deep Learning-Enabled Automated Quality Control for Liver MR Elastography: Initial Results,” was published in the Journal of Magnetic Resonance Imaging.
Using five deep learning training models, an accuracy of 92% was achieved by the best-performing ensemble on retrospective MRE images of patients with varied liver stiffnesses. The team also achieved a return of the analyzed data within seconds. The rapidity of image quality return allows the technician to focus on adjusting hardware or patient orientation for re-scan in a single session, rather than requiring patients to return for costly and timely re-scans due to low-quality initial images.
This new research is a step toward streamlining the review pipeline for MRE using deep learning techniques, which have remained unexplored compared to other medical imaging modalities. The research also provides a helpful baseline for future avenues of inquiry, such as assessing the health of the spleen or kidneys. It may also be applied to automation for image quality control for monitoring non-hepatic conditions, such as breast cancer or muscular dystrophy, in which tissue stiffness is an indicator of initial health and disease progression. Ueda, Nieves, and their team hope to test these models on Siemens Healthineers magnetic resonance scanners within the next year.
Publication
Nieves-Vazquez, H.A., Ozkaya, E., Meinhold, W., Geahchan, A., Bane, O., Ueda, J. and Taouli, B. (2024), Deep Learning-Enabled Automated Quality Control for Liver MR Elastography: Initial Results. J Magn Reson Imaging. https://doi.org/10.1002/jmri.29490
Prior Work
Robotically Precise Diagnostics and Therapeutics for Degenerative Disc Disorder
Related Material
Editorial for “Deep Learning-Enabled Automated Quality Control for Liver MR Elastography: Initial Results”
News Contact
Christa M. Ernst |
Research Communications Program Manager |
Topic Expertise: Robotics, Data Sciences, Semiconductor Design & Fab |
Jul. 11, 2024
Georgia Tech School of Electrical and Computer Engineering (ECE) Professor Vince Calhoun was honored with the Organization for Human Brain Mapping’s Glass Brain Award at the organization’s annual meeting in Seoul, Korea, this June.
The award recognizes lifetime achievements by leading researchers using or facilitating neuroimaging to discover original and influential findings regarding the organization and function of the human brain.
“One of the things I really like about the Glass Brain is it symbolizes to me our desire to see beneath the surface. It's a challenge to be open and transparent in our work,” Calhoun said. “Visualization is hard and subjective, we have to work to make the links between our fancy models, the underlying data, and our conclusions about the brain, as transparent as possible.”
He's made many contributions to the field, including authorship on over 1,000 papers in industry journals, generating over 100,000 citations within the research community.
Much of his work has focused on developing data-driven approaches to study brain dynamics, multimodal neuroimaging of the brain, and brain biomarkers across a range of mental and neurological conditions.
Calhoun is the founding director of the tri-institutional Center for Translational Research in Neuroimaging and Data Science (TReNDS), a joint effort between Georgia State, Georgia Tech, and Emory University. The Center’s goal is to improve the understanding of the human brain using advanced analytic approaches with an emphasis on translational research such as the development of predictive biomarkers for mental and neurological disorders.
Calhoun received his bachelor’s degree in electrical engineering (EE) from the University of Kansas, a pair of master’s degrees in biomedical engineering and information systems from Johns Hopkins University, and a Ph.D. in EE from the University of Maryland Baltimore County.
Before coming to Georgia Tech, he worked as a research engineer in the psychiatric neuroimaging laboratory at Johns Hopkins, the director of medical image analysis at the Olin Neuropsychiatry Research Center. He’s also been an associate professor at Yale University, a distinguished professor at the University of New Mexico, and the President of the Mind Research Network.
The Glass Brain Award is the latest of many distinctions Calhoun has received during his accomplished career. He’s received fellowships from a number of organizations, including the Institute of Electrical and Electronic Engineers (IEEE), the Association for the Advancement of Science, and the American Institute of Biomedical and Medical Engineers.
He’s also won an IEEE Outstanding Engineer Award in 2014 and an IEEE Southwest Area Outstanding Educator Award in 2015.
News Contact
Zachary Winiecki
Jul. 09, 2024
From plaque sticking to teeth to scum on a pond, biofilms can be found nearly everywhere. These colonies of bacteria grow on implanted medical devices, our skin, contact lenses, and in our guts and lungs. They can be found in sewers and drainage systems, on the surface of plants, and even in the ocean.
“Some research says that 80% of infections in human bodies can be attributed to the bacteria growing in biofilms,” Aawaz Pokhrel says, lead author of a groundbreaking new study that uses physics to investigate how these biofilms grow.
The paper, “The Biophysical Basis of Bacterial Colony Growth,” was published in Nature Physics this week, and it shows that the fitness of a biofilm — its ability to grow, expand, and absorb nutrients from the medium or the substrate — is largely impacted by the contact angle that the biofilm’s edge makes with the substrate. The study also found that this geometry has a bigger influence on fitness than anything else, including the rate at which the cells can reproduce.
“That was the big surprise for us,” says corresponding author Peter Yunker, an associate professor in Georgia Tech’s School of Physics. “We expected that the geometry would play an important role, and we thought that figuring out exactly what the geometry is would be important for understanding why the range expansion rate, for example, [the rate at which the biofilm spreads across the surface over time] is constant. But we didn't start the project thinking that geometry would be the single most important factor.”
Understanding how biofilms grow — and what factors contribute to their growth rate — could lead to critical insights on controlling them, with applications for human health, like slowing the spread of infection or creating cleaner surfaces. “What got me excited was this opportunity to use physics to learn about complex biological systems,” Pokhrel, who is also a Ph.D. student in Yunker’s lab, adds. “Especially on a project that has so many applications. The combination of the importance for human health and exciting research was really intriguing for me.”
A new method
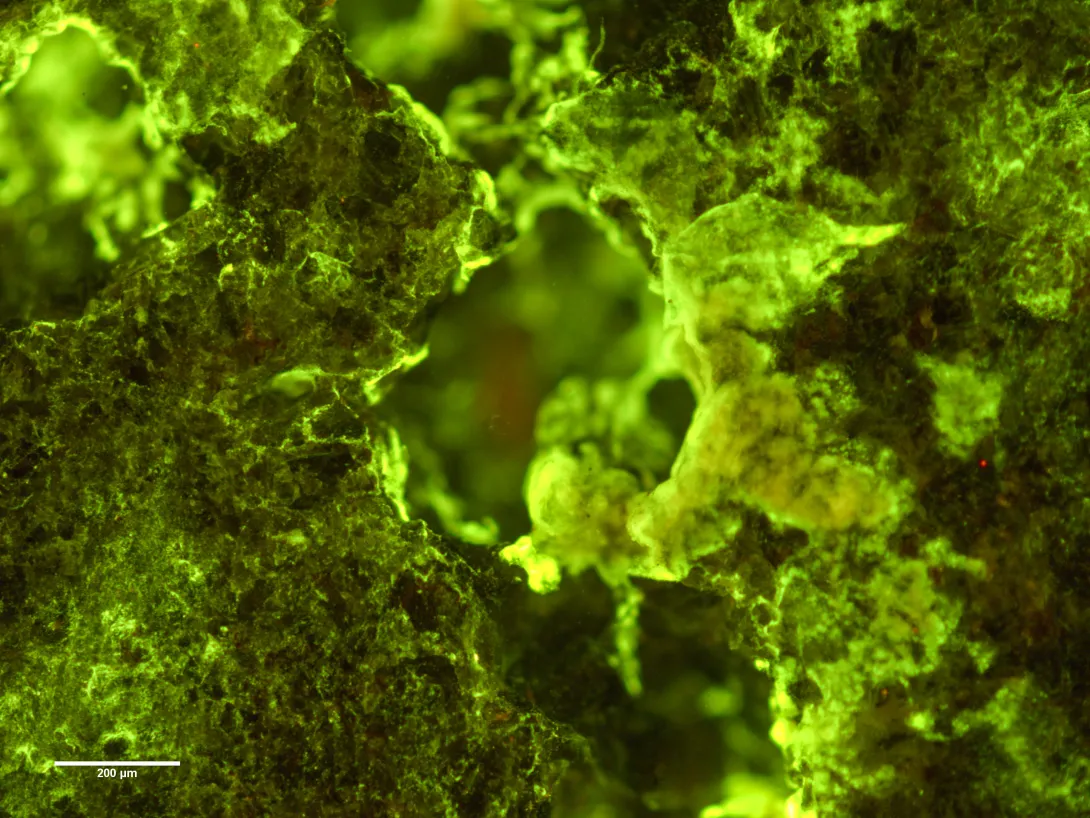
While biofilms are ubiquitous in nature, studying them has proven difficult. Because these “cities of microorganisms” are comprised of tiny individuals, scientists have struggled to image them successfully.
That changed in 2015, when Yunker began wondering if interferometry, a commonly used imaging technique in physics and materials science, could be applied to biofilms. “Given my background in physics, I was familiar with its use in materials applications,” Yunker recalls. “I thought applying this technique more broadly might be interesting, because we know from decades of physics that surface interfaces contain a lot of information about the processes that create them.”
The technique proved to be simple, effective, and time-efficient, providing nanometer-scale resolution of bacterial colonies. “It allows us to essentially get a picture of the topography — the shape of the surface of the bacterial population — with super-resolution,” Yunker adds.
Leveraging interferometry, the team began conducting new biofilm experiments, investigating how colonies’ shapes changed over time. Co-first author Gabi Steinbach, formerly a postdoctoral scholar in Yunker’s lab and now a scientific research coordinator at the University of Maryland, noticed that every colony had a specific shape when it was small: a spherical cap, like a slice from the top of a sphere, or a droplet of water. It’s a shape that shows up often in physics, and that sparked the team’s interest.
“A spherical cap in physics is very interesting, because it is a surface-minimizing shape,” Pokhrel adds. “I was curious why a biological material was growing in this shape, and we started wondering if there was some physics to it – perhaps geometry was involved. And that made us think that maybe we could develop a model. And that got me really excited.”
A mathematical mystery
However, the researchers soon hit a roadblock. “While we could see that the colonies were spherical caps at first, they would deviate from that shape as they grew,” Pokhrel says. “And the shape that they grew into was difficult to describe with existing spherical cap geometry.”
“The middle didn’t grow as quickly as it should to keep the spherical cap shape, and we wanted to connect all of this to the range expansion [the rate at which the colony spread across a surface],” Yunker adds. “But we knew that somehow, geometry was playing a very important role.”
Finally, Thomas Day, a former graduate student in Yunker’s lab, now a postdoctoral fellow at the University of Southern California, and one of the authors of the paper, suggested a quirky problem of geometry called the napkin ring problem.
“As soon as we started to think about the napkin ring problem, we were able to start developing a mathematical toolkit,” Yunker says, though the solution wasn’t effortless. “We couldn't find anyone who had ever looked at a spherical cap napkin ring before, because the application is very rare.”
Pokhrel, alongside two co-authors, was responsible for working out the geometry. He discovered that the cells grew exponentially at the edge of the shape, expanding further onto the medium, while the cells in the middle grew upward, creating a shape not unlike an egg in a frying pan — if the egg white was expanding outwards, while the yolk was only growing taller.
This was the breakthrough discovery: Because the cells at the middle were only contributing to the biofilm’s height, the team only needed to account for how many cells were at the edge of the biofilm, and the shape they needed to be in to grow and spread.
After incorporating their findings into a mathematical model, the team found that the contact angle was the most important factor: the angle that the very edge of the biofilm made when it touched the surface it was growing on. That single geometric quality is even more important to a biofilm’s growth than the rate at which it can reproduce cells.
The physics-biology connection
Overall, the project took more than three years, from conception to publication. “Aawaz really made an incredible effort seeing this work through,” Yunker says. “It was many years and many, many experiments. But the finished product is 100% worth it.”
The team hopes the research will pave the way for future studies, which could lead to applications like controlling biofilm growth to help prevent infections.
“Going forward, there are still a lot of research avenues,” Pokhrel says. “For example, looking at competition experiments between biofilms — do taller colonies change their contact angle so that they can spread faster? What role does this geometry play in competition?”
“Biology is complex,” Yunker adds. In nature, the surface a biofilm grows on may not be as consistent as a laboratory surface, and colonies may have different mutations or may consist of more than one species. And while the model is based on how biofilms behave in a controlled lab environment, it’s a critical first step in understanding how they may behave in nature.
Citation: Pokhrel, A.R., Steinbach, G., Krueger, A. et al. The biophysical basis of bacterial colony growth. Nat. Phys. (2024). https://doi.org/10.1038/s41567-024-02572-3
Funding information: This research was funded by the NIH National Institute of General Medical Sciences and NSF Biomaterials
News Contact
Written by Selena Langner
Jun. 26, 2024
Georgia Tech School of Electrical and Computer Engineering (ECE) second-year Ph.D. student Aref Valipour was awarded a GT Next Award for his MicroCodes technology, which shows utility in aiding with cellular characterization and management.
The award is given annually by the Office of Technology Licensing (OTL), recognizing the inventiveness of Georgia Tech's graduate students and postdocs. Winners submit projects that show commercial promise and a commitment to research and development that can contribute significantly to the betterment of society.
MicroCodes is comprised of a new microfabrication method and coding method. Initially developed as a facilitator in the characterization of Circulating Tumor Cells (CTCs), it later evolved to become a universal add-on for a number of other medical and MEMS applications.
The award acts as a direct financial investment into the future of the technology, helping to carry out different commercial processes, such as procuring materials and supplies, and carrying out customer discovery initiatives.
Valipour was one of five winners this year.
His research interests are centered around MEMS, microfabrication, bioelectronics and utilizing micro and nano engineering to understand how cancer works and how it can be managed.
He’s developed technologies and methods to characterize, monitor, and intervene in cancer to ultimately create the platform that gives an edge in the fight against cancer. To date, these efforts have resulted in seven invention disclosures to OTL, with the award-winning MicroCodes being the latest.
Valipour is currently a member of Biomedical Microsystems Laboratory. Outside of ECE, he is part of the Technology Innovation: Generating Economic Results (TI:GER) program at Scheller College of Business.
News Contact
Zachary Winiecki
Jun. 24, 2024
The Children's Healthcare of Atlanta Pediatric Technology Center at Georgia Tech (PTC) is excited to announce that Gian-Gabriel Garcia will serve as its Pillar 1 Co-Lead. Pillar 1 focuses on data science, machine learning, and artificial intelligence. In his new role, Garcia’s responsibilities will include setting the pillar’s strategy and vision, selecting and managing projects, overseeing various pillar activities, and working collaboratively across research groups and institutions. He will also identify cutting-edge technology and engineering solutions to implement priority projects while balancing the pragmatism and feasibility of these approaches.
The PTC brings clinical experts together with Georgia Tech scientists and engineers to develop technological solutions to problems in the health and care of children. The Center provides extraordinary opportunities for interdisciplinary collaboration in pediatrics, creating breakthrough discoveries that often can only be found at the intersection of multiple disciplines.
Garcia will work under the leadership of PTC Co-Directors Dr. Stanislav Emelianov (Georgia Tech) and Dr. Wilbur Lam (Children’s) of Georgia Tech’s Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. Dr. Naveen Muthu of Children’s Physician Group will be Garcia’s counterpart in leading Pillar 1.
Since 2021, Garcia has served as an assistant professor in Georgia Tech’s H. Milton Stewart School of Industrial and Systems Engineering. His research group has published numerous journal and conference papers, and book chapters related to data-driven machine learning and optimization in healthcare, including various applications in diagnosis and disease management of concussion, opioids, cardiovascular disease, glaucoma, and maternal health. He has received federal funding as a primary investigator from both the National Institutes for Health and the Agency for Healthcare Research and Quality. He and his research group have received several national and international recognitions for their work.
Garcia also teaches graduate-level courses in machine learning and optimization for healthcare. He received his Ph.D. in industrial and operations engineering at the University of Michigan and was a postdoctoral fellow at the MGH Institute for Technology Assessment.
Jun. 12, 2024
Adoptive T-cell therapy has revolutionized medicine. A patient’s T-cells — a type of white blood cell that is part of the body’s immune system — are extracted and modified in a lab and then infused back into the body, to seek and destroy infection, or cancer cells.
Now Georgia Tech bioengineer Ankur Singh and his research team have developed a method to improve this pioneering immunotherapy.
Their solution involves using nanowires to deliver therapeutic miRNA to T-cells. This new modification process retains the cells’ naïve state, which means they’ll be even better disease fighters when they’re infused back into a patient.
“By delivering miRNA in naïve T cells, we have basically prepared an infantry, ready to deploy,” Singh said. “And when these naïve cells are stimulated and activated in the presence of disease, it’s like they’ve been converted into samurais.”
Lean and Mean
Currently in adoptive T-cell therapy, the cells become stimulated and preactivated in the lab when they are modified, losing their naïve state. Singh’s new technique overcomes this limitation. The approach is described in a new study published in the journal Nature Nanotechnology.
“Naïve T-cells are more useful for immunotherapy because they have not yet been preactivated, which means they can be more easily manipulated to adopt desired therapeutic functions,” said Singh, the Carl Ring Family Professor in the Woodruff School of Mechanical Engineering and the Wallace H. Coulter Department of Biomedical Engineering.
The raw recruits of the immune system, naïve T-cells are white blood cells that haven’t been tested in battle yet. But these cellular recruits are robust, impressionable, and adaptable — ready and eager for programming.
“This process creates a well-programmed naïve T-cell ideal for enhancing immune responses against specific targets, such as tumors or pathogens,” said Singh.
The precise programming naïve T-cells receive sets the foundational stage for a more successful disease fighting future, as compared to preactivated cells.
Giving Fighter Cells a Boost
Within the body, naïve T-cells become activated when they receive a danger signal from antigens, which are part of disease-causing pathogens, but they send a signal to T-cells that activate the immune system.
Adoptive T-cell therapy is used against aggressive diseases that overwhelm the body’s defense system. Scientists give the patient’s T-cells a therapeutic boost in the lab, loading them up with additional medicine and chemically preactivating them.
That’s when the cells lose their naïve state. When infused back into the patient, these modified T-cells are an effective infantry against disease — but they are prone to becoming exhausted. They aren’t samurai. Naïve T-cells, though, being the young, programmable recruits that they are, could be.
The question for Singh and his team was: How do we give cells that therapeutic boost without preactivating them, thereby losing that pristine, highly suggestable naïve state? Their answer: Nanowires.
NanoPrecision: The Pointed Solution
Singh wanted to enhance naïve T-cells with a dose of miRNA. miRNA is a molecule that, when used as a therapeutic, works as a kind of volume knob for genes, turning their activity up or down to keep infection and cancer in check. The miRNA for this study was developed in part by the study’s co-author, Andrew Grimson of Cornell University.
“If we could find a way to forcibly enter the cells without damaging them, we could achieve our goal to deliver the miRNA into naïve T cells without preactivating them,” Singh explained.
Traditional modification in the lab involves binding immune receptors to T-cells, enabling the uptake of miRNA or any genetic material (which results in loss of the naïve state). “But nanowires do not engage receptors and thus do not activate cells, so they retain their naïve state,” Singh said.
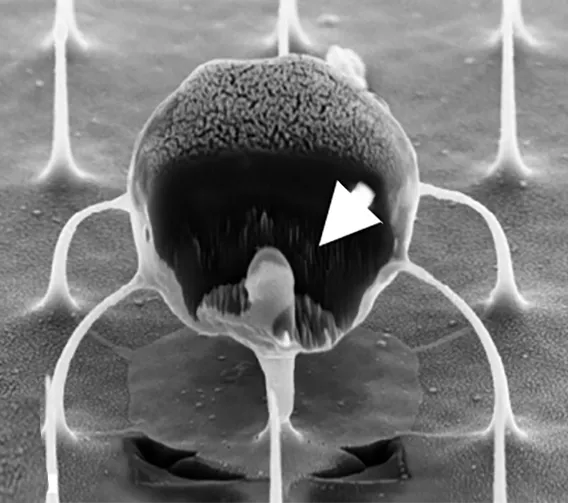
The nanowires, silicon wafers made with specialized tools at Georgia Tech’s Institute for Electronics and Nanotechnology, form a fine needle bed. Cells are placed on the nanowires, which easily penetrate the cells and deliver their miRNA over several hours. Then the cells with miRNA are flushed out from the tops of the nanowires, activated, eventually infused back into the patient. These programmed cells can kill enemies efficiently over an extended time period.
“We believe this approach will be a real gamechanger for adoptive immunotherapies, because we now have the ability to produce T-cells with predictable fates,” says Brian Rudd, a professor of immunology at Cornell University, and co-senior author of the study with Singh.
The researchers tested their work in two separate infectious disease animal models at Cornell for this study, and Singh described the results as “a robust performance in infection control.”
In the next phase of study, the researchers will up the ante, moving from infectious disease to test their cellular super soldiers against cancer and move toward translation to the clinical setting. New funding from the Georgia Clinical & Translational Science Alliance is supporting Singh’s research.
CITATION: Kristel J. Yee Mon, Sungwoong Kim, Zhonghao Dai, Jessica D. West, Hongya Zhu5, Ritika Jain, Andrew Grimson, Brian D. Rudd, Ankur Singh. “Functionalized nanowires for miRNA-mediated therapeutic programming of naïve T cells,” Nature Nanotechnology.
FUNDING: Curci Foundation, NSF (EEC-1648035, ECCS-2025462, ECCS-1542081), NIH (5R01AI132738-06, 1R01CA266052-01, 1R01CA238745-01A1, U01CA280984-01, R01AI110613 and U01AI131348).
News Contact
Jerry Grillo
May. 23, 2024
Georgia Tech School of Electrical and Computer Engineering (ECE) Ph.D. candidate Jiaao Lu has been awarded a Verbal Poster Award at InterfaceRice 2024, which was held in Houston, Texas from April 30 to May 1.
The research, done in collaboration with Emory University and Johns Hopkins University, focuses on new device technologies to support innovation in optogenetics, a technique used to control neuron activity or other cell types with light.
The two-day conference, hosted by Rice University and the Texas Medical Center, connects researchers, clinicians, and business leaders in the fields of neuroengineering, neurotechnology, neuroscience, and neurosurgery.
The award recognizes excellence in oral presentation among presented posters. There were 70 total posters accepted to this year’s conference.
Lu and fellow researchers in ECE Professor Muhannad Bakir’s Integrated 3D System Group are working on integrating a μLED chip with a flexible microelectrode array (MEA) for optogenetic activation, while simultaneously recording electrical activity in muscle tissue.
Recently, they developed an MEA with a single blue LED, which allows for targeted activation of specific opsins. Opsins are light-sensitive proteins that can be expressed in specific cells to control their activity using light.
So far, the technology has been successful in activating facial muscles and inducing jaw movement in transgenic mice.
“I had a rewarding time at the InterfaceRice conference,” Lu said. “The feedback on my project was highly encouraging, and discussions extended beyond engineers to include clinicians - actual users of my technology - who offered invaluable insights. These conversations also led to several opportunities for potential collaborations, which was very exciting.”
The next step in the research for Lu and her collaborators is to add multiple optical channels and incorporate more LED colors, allowing them to activate and observe more opsins.
News Contact
Zachary Winiecki
May. 21, 2024
Asthma impacts more than 40 million Americans, and 10% of the world’s population. However, current anti-inflammatory treatments only partially control the disease’s symptoms. Now, Liang Han, an associate professor in the School of Biological Sciences, has been awarded a $2.47M grant by the National Institute of Health to study the role our nervous system plays in asthma — and the potential for new treatments. The grant will fund five years of research, with work beginning this spring.
“Asthma is typically considered an allergic inflammatory disease,” Han says, “and so the majority of research has previously focused on immune responses. But there is emerging evidence that the nervous system plays a critical role in the disease.”
Han highlights that our lungs are full of sensory nerves, which help monitor their internal state, and play an important role in regulating our breathing patterns and respiratory system. Vagal sensory neurons help send information from the lungs to the brain. Recent data collected by Yanyan Xing, a former postdoctoral researcher in the Han lab and now a scientist at Empress Therapeutics, suggested that blocking a group of vagal sensory neurons stopped the development of asthma symptoms in mice.
“Since these sensory neurons are responsible for responses like coughing, bronchoconstriction, and mucus secretion, all of which are asthma symptoms, we want to investigate whether blocking these neurons can help inhibit asthma in humans,” Han says. “If so, this might prove a promising treatment avenue for asthma.”
The nervous system connection
In her lab at Georgia Tech, Han’s research team investigates the role the nervous system plays in creating and behavioral responses, and how that contributes to chronic diseases. “We want to understand how the nervous system receives, transmits, and interprets various stimuli to induce physiological and behavioral responses,” she explains.
This year, Han also received a $550k grant from the National Science Foundation to investigate the neural circuit controlling itch sensation. The research has the potential to uncover new treatments for sensory conditions like chronic itch.
News Contact
Written by Selena Langner
May. 15, 2024
Georgia Tech researchers say non-English speakers shouldn’t rely on chatbots like ChatGPT to provide valuable healthcare advice.
A team of researchers from the College of Computing at Georgia Tech has developed a framework for assessing the capabilities of large language models (LLMs).
Ph.D. students Mohit Chandra and Yiqiao (Ahren) Jin are the co-lead authors of the paper Better to Ask in English: Cross-Lingual Evaluation of Large Language Models for Healthcare Queries.
Their paper’s findings reveal a gap between LLMs and their ability to answer health-related questions. Chandra and Jin point out the limitations of LLMs for users and developers but also highlight their potential.
Their XLingEval framework cautions non-English speakers from using chatbots as alternatives to doctors for advice. However, models can improve by deepening the data pool with multilingual source material such as their proposed XLingHealth benchmark.
“For users, our research supports what ChatGPT’s website already states: chatbots make a lot of mistakes, so we should not rely on them for critical decision-making or for information that requires high accuracy,” Jin said.
“Since we observed this language disparity in their performance, LLM developers should focus on improving accuracy, correctness, consistency, and reliability in other languages,” Jin said.
Using XLingEval, the researchers found chatbots are less accurate in Spanish, Chinese, and Hindi compared to English. By focusing on correctness, consistency, and verifiability, they discovered:
- Correctness decreased by 18% when the same questions were asked in Spanish, Chinese, and Hindi.
- Answers in non-English were 29% less consistent than their English counterparts.
- Non-English responses were 13% overall less verifiable.
XLingHealth contains question-answer pairs that chatbots can reference, which the group hopes will spark improvement within LLMs.
The HealthQA dataset uses specialized healthcare articles from the popular healthcare website Patient. It includes 1,134 health-related question-answer pairs as excerpts from original articles.
LiveQA is a second dataset containing 246 question-answer pairs constructed from frequently asked questions (FAQs) platforms associated with the U.S. National Institutes of Health (NIH).
For drug-related questions, the group built a MedicationQA component. This dataset contains 690 questions extracted from anonymous consumer queries submitted to MedlinePlus. The answers are sourced from medical references, such as MedlinePlus and DailyMed.
In their tests, the researchers asked over 2,000 medical-related questions to ChatGPT-3.5 and MedAlpaca. MedAlpaca is a healthcare question-answer chatbot trained in medical literature. Yet, more than 67% of its responses to non-English questions were irrelevant or contradictory.
“We see far worse performance in the case of MedAlpaca than ChatGPT,” Chandra said.
“The majority of the data for MedAlpaca is in English, so it struggled to answer queries in non-English languages. GPT also struggled, but it performed much better than MedAlpaca because it had some sort of training data in other languages.”
Ph.D. student Gaurav Verma and postdoctoral researcher Yibo Hu co-authored the paper.
Jin and Verma study under Srijan Kumar, an assistant professor in the School of Computational Science and Engineering, and Hu is a postdoc in Kumar’s lab. Chandra is advised by Munmun De Choudhury, an associate professor in the School of Interactive Computing.
The team will present their paper at The Web Conference, occurring May 13-17 in Singapore. The annual conference focuses on the future direction of the internet. The group’s presentation is a complimentary match, considering the conference's location.
English and Chinese are the most common languages in Singapore. The group tested Spanish, Chinese, and Hindi because they are the world’s most spoken languages after English. Personal curiosity and background played a part in inspiring the study.
“ChatGPT was very popular when it launched in 2022, especially for us computer science students who are always exploring new technology,” said Jin. “Non-native English speakers, like Mohit and I, noticed early on that chatbots underperformed in our native languages.”
School of Interactive Computing communications officer Nathan Deen and School of Computational Science and Engineering communications officer Bryant Wine contributed to this report.
News Contact
Bryant Wine, Communications Officer
bryant.wine@cc.gatech.edu
Nathan Deen, Communications Officer
ndeen6@cc.gatech.edu
May. 14, 2024
The call from his mom is still vivid 20 years later. Moments this big and this devastating can define lives, and for Hong Yeo, today a Georgia Tech mechanical engineer, this call certainly did. Yeo was a 21-year-old in college studying car design when his mom called to tell him his father had died in his sleep. A heart attack claimed the life of the 49-year-old high school English teacher who had no history of heart trouble and no signs of his growing health threat. For the family, it was a crushing blow that altered each of their paths.
“It was an uncertain time for all of us,” said Yeo. “This loss changed my focus.”
For Yeo, thoughts and dreams of designing cars for Hyundai in Korea turned instead toward medicine. The shock of his father going from no signs of illness to gone forever developed into a quest for medical answers that might keep other families from experiencing the pain and loss his family did — or at least making it less likely to happen.
Yeo’s own research and schooling in college pointed out a big problem when it comes to issues with sleep and how our bodies’ systems perform — data. He became determined to invent a way to give medical doctors better information that would allow them to spot a problem like his father’s before it became life-threatening.
His answer: a type of wearable sleep data system. Now very close to being commercially available, Yeo’s device comes after years of working on the materials and electronics for an easy-to-wear, comfortable mask that can gather data about sleep over multiple days or even weeks, allowing doctors to catch sporadic heart problems or other issues. Different from some of the bulky devices with straps and cords currently available for at-home heart monitoring, it offers the bonuses of ease of use and comfort, ensuring little to no alteration to users’ bedtime routine or wear. This means researchers can collect data from sleep patterns that are as close to normal sleep as possible.
“Most of the time now, gathering sleep data means the patient must come to a lab or hospital for sleep monitoring. Of course, it’s less comfortable than home, and the devices patients must wear make it even less so. Also, the process is expensive, so it’s rare to get multiple nights of data,” says Audrey Duarte, University of Texas human memory researcher.
Duarte has been working with Yeo on this system for more than 10 years. She says there are so many mental and physical health outcomes tied to sleep that good, long-term data has the potential to have tremendous impact.
“The results we’ve seen are incredibly encouraging, related to many things —from heart issues to areas I study more closely like memory and Alzheimer’s,” said Duarte.
Yeo’s device may not have caught the arrhythmia that caused his father’s heart attack, but nights or weeks of data would have made effective medical intervention much more likely.
Inspired by his own family’s loss, Yeo’s life’s work has become a tool of hope for others.
Pagination
- Previous page
- Page 19
- Next page