Oct. 28, 2024
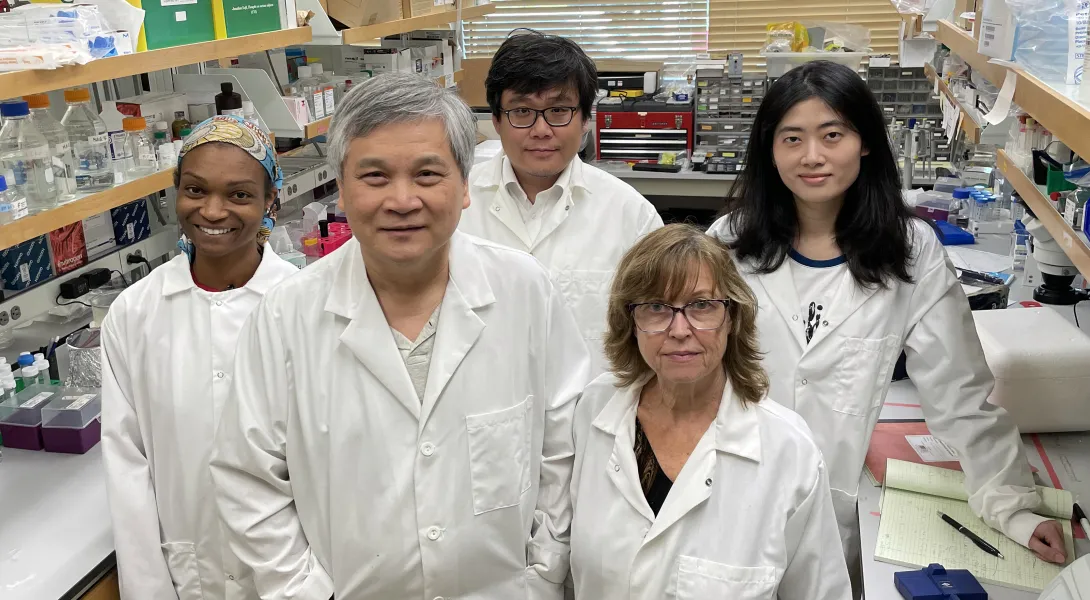
Simone Douglas-Green, an asst. professor at the Wallace H. Coulter Department of Biomedical Engineering, has recently been awarded the 2024 Sloan Scholars Mentoring Network (SSMN) seed grant.
This $10,000 award, the first independent funding for the Douglas-Green lab, will support her research on protein coronas — protein complexes that form when proteins are absorbed to the surface of nanoparticles.
“We’re excited about receiving the grant,” said Douglas-Green. “Sloan was integral to my career as a Ph.D. student, and now the SSMN Seed Grant is a catalyst for our research group’s first project.”
The grant was established specifically to support research. But the SSMN program, formed through a collaboration between the Social Science Research Council and the Alfred P. Sloan Foundation, does more than offer grants. It also provides workshops, mentoring, and networking opportunities to support scholars in their academic journey.
With this grant, the Douglas-Green lab will address the challenge of isolating and characterizing protein coronas on nanoparticles, particularly small ones like polyamidoamine (PAMAM) dendrimers, a class of nanoparticles which hold significant potential for drug delivery.
“Having the right tools and techniques to accurately study bio-nano interactions will make this endeavor possible, and our new group will be at the forefront in developing them,” said Douglas-Green.
What sets her research apart is her lab’s use of the electrophoresis process to characterize protein coronas. Building on her post-doctoral research at the Massachusetts Institute of Technology (MIT), Douglas-Green’s lab is employing a nondenaturing electrophoresis technique combined with mass spectrometry to separate and identify these tiny protein coronas.
Electrophoresis is a lab technique used to separate molecules, like proteins, based on their size and charge, helping scientists understand the composition and behavior of these proteins. When an electric current is applied, proteins move through a gel at different speeds depending on their size and charge, so researchers can analyze the types of proteins involved. The technique can be used to study nanoparticle-protein complexes on nanoparticles with smaller sizes and densities in ways that existing methods, like centrifugation or dynamic light scattering, cannot.
Douglas-Green’s lab has turned the challenges of nanoparticle-protein separation into an opportunity for innovation. By avoiding the use of SDS, a common detergent that interferes with PEG (a polymer that makes nanoparticles more compatible with the body), they found a way to better characterize protein coronas on PEG-coated nanocarriers.
The separation technique developed by Douglas-Green’s lab ensured compatibility with the surface chemistry of dendrimers, a priority as the research team continues advancing nanoparticle separation techniques.
“Our engineering goal is to develop tools and techniques to study protein coronas on various types of nanoparticles,” said Douglas-Green. “Using these tools, we can uncover some biological questions to understand person- or disease-specific protein coronas.”
The work is part of a broader effort to build a toolkit of techniques for scientists to better understand bio-nanoparticles interactions–leading to more targeted nanoparticle-based treatments and therapeutics.
“For me, this is more than seed funding to pursue science,” Douglas-Green said. “This is funding the start of a dream realized!”
News Contact
By Yanet Chernet
Contact:
Kelly Petty
Communications
Wallace H. Coulter Department of Biomedical Engineering
Nov. 19, 2024
Three College of Sciences students with aspirations of making a difference in medicine were selected as recipients of the prestigious Stamps President’s Scholarship. Though this scholarship is typically given to 40 exceptional incoming first-year students, a select few second- and third-year students are chosen to receive the honor for exemplifying the program’s pillars of scholarship, leadership, progress, and service.
The new Scholars include School of Biological Sciences/School of Modern Languages student Sonali Kaluri, School of Chemistry and Biochemistry student Seth Kinoshita, and School of Biological Sciences student Medina McCowin.
As part of the program, the selected students will receive a full-ride scholarship, special mentoring, and travel opportunities.
About the Scholars
Sonali Kaluri is a third-year student double majoring in biology and applied languages and intercultural studies (with a concentration in Spanish). Deeply passionate about women's health, she has researched clinical considerations of treating liver disease in pregnant women and the impact of a virtual lactation program on maternal and infant health outcomes at the University of Massachusetts Medical School. In her spare time, she volunteers at the Winship Cancer Institute and the March of Dimes and is a member of the Yellow Jacket Fencing Club.
“I hope to attend medical school and pursue a career in academic medicine after graduation from Georgia Tech,” says Kaluri. “My research experience has made me acutely aware of the gaps in medical knowledge regarding the different ways disease processes affect women, and I hope to become an advocate for change through research and clinical practice!”
Seth Kinoshita is a third-year biochemistry major with a minor in health and medical sciences. As an undergraduate research assistant with the Department of Biomedical Engineering, he focuses on a novel drug delivery structure that can be surgically inserted to decrease recovery time and minimize invasiveness for tendon injuries. His work has been published in several academic journals. He serves as an undergraduate research ambassador and a pre-health mentor — and spends his free time with Sympathetic Vibrations, Georgia Tech's male a cappella group. Kinoshita also works as the medical coordinator for Aurora Day Camp, a camp for children with cancer and their siblings.
"After graduation, I want to pursue an M.D./Ph.D. in regenerative orthopedic medicine to bridge my tendon repair research with direct implementation into patients,” says Kinoshita. “I aim to develop innovative treatments that can restore mobility in the extremities and improve the quality of life for patients with musculoskeletal disorders."
Medina McCowin is a third-year biology major researching cancer treatment methods in the Sulchek BioMEMS and Biomechanics Lab. She also worked for Lachance Laboratories as an undergraduate researcher, investigating cancer genetics. Active on campus, she is the biology representative for the Georgia Tech Undergraduate House of Representatives and president of the Georgia Tech Public Health Student Association. McCowin has also held several leadership roles with the Georgia Tech American Medical Student Association.
“In the future, I hope to pursue an M.D./Ph.D. and become a pediatric oncologist and cancer treatment researcher, focusing on improving pediatric cancer treatments,” says McCowin. “Working in the healthcare field and experiencing personal loss has taught me that empathy and compassion are the most important factors in becoming a doctor. As a doctor, I want to contribute to the advancements of pediatric medicine, but also be dedicated to improving the emotional and mental well-being of my patients and their families.”
News Contact
Writer: Laura S. Smith
College of Sciences
laura.smith@cos.gatech.edu
Nov. 21, 2024
Amino acids are essential for nearly every process in the human body. Often referred to as ‘the building blocks of life,’ they are also critical for commercial use in products ranging from pharmaceuticals and dietary supplements, to cosmetics, animal feed, and industrial chemicals.
And while our bodies naturally make amino acids, manufacturing them for commercial use can be costly — and that process often emits greenhouse gasses like carbon dioxide (CO2).
In a landmark study, a team of researchers has created a first-of-its kind methodology for synthesizing amino acids that uses more carbon than it emits. The research also makes strides toward making the system cost-effective and scalable for commercial use.
“To our knowledge, it’s the first time anyone has synthesized amino acids in a carbon-negative way using this type of biocatalyst,” says lead corresponding author Pamela Peralta-Yahya, who emphasizes that the system provides a win-win for industry and environment. “Carbon dioxide is readily available, so it is a low-cost feedstock — and the system has the added bonus of removing a powerful greenhouse gas from the atmosphere, making the synthesis of amino acids environmentally friendly, too.”
The study, “Carbon Negative Synthesis of Amino Acids Using a Cell-Free-Based Biocatalyst,” published today in ACS Synthetic Biology, is publicly available. The research was led by Georgia Tech in collaboration with the University of Washington, Pacific Northwest National Laboratory, and the University of Minnesota.
The Georgia Tech research contingent includes Peralta-Yahya, a professor with joint appointments in the School of Chemistry and Biochemistry and School of Chemical and Biomolecular Engineering (ChBE); first author Shaafique Chowdhury, a Ph.D. student in ChBE; Ray Westenberg, a Ph.D student in Bioengineering; and Georgia Tech alum Kimberly Wennerholm (B.S. ChBE ’23).
Costly chemicals
There are two key challenges to synthesizing amino acids on a large scale: the cost of materials, and the speed at which the system can generate amino acids.
While many living systems like cyanobacteria can synthesize amino acids from CO2, the rate at which they do it is too slow to be harnessed for industrial applications, and these systems can only synthesize a limited number of chemicals.
Currently, most commercial amino acids are made using bioengineered microbes. “These specially designed organisms convert sugar or plant biomass into fuel and chemicals,” explains first author Chowdhury, “but valuable food resources are consumed if sugar is used as the feedstock — and pre-processing plant biomass is costly.” These processes also release CO2 as a byproduct.
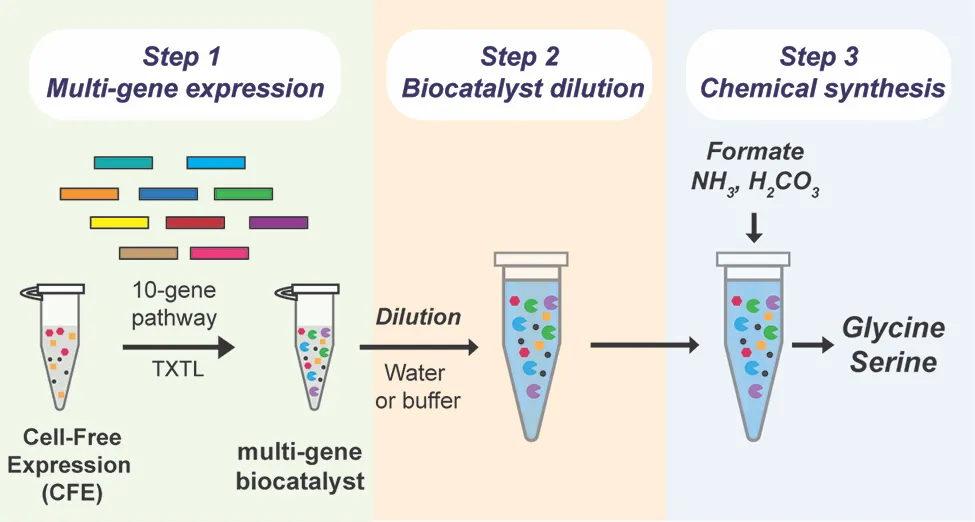
Chowdhury says the team was curious “if we could develop a commercially viable system that could use carbon dioxide as a feedstock. We wanted to build a system that could quickly and efficiently convert CO2 into critical amino acids, like glycine and serine.”
The team was particularly interested in what could be accomplished by a ‘cell-free’ system that leveraged some process of a cellular system — but didn’t actually involve living cells, Peralta-Yahya says, adding that systems using living cells need to use part of their CO2 to fuel their own metabolic processes, including cell growth, and have not yet produced sufficient quantities of amino acids.
“Part of what makes a cell-free system so efficient,” Westenberg explains, “is that it can use cellular enzymes without needing the cells themselves. By generating the enzymes and combining them in the lab, the system can directly convert carbon dioxide into the desired chemicals. Because there are no cells involved, it doesn’t need to use the carbon to support cell growth — which vastly increases the amount of amino acids the system can produce.”
A novel solution
While scientists have used cell-free systems before, one of the necessary chemicals, the cell lysate biocatalyst, is extremely costly. For a cell-free system to be economically viable at scale, the team needed to limit the amount of cell lysate the system needed.
After creating the ten enzymes necessary for the reaction, the team attempted to dilute the biocatalyst using a technique called ‘volumetric expansion.’ “We found that the biocatalyst we used was active even after being diluted 200-fold,” Peralta-Yahya explains. “This allows us to use significantly less of this high-cost material — while simultaneously increasing feedstock loading and amino acid output.”
It’s a novel application of a cell-free system, and one with the potential to transform both how amino acids are produced, and the industry’s impact on our changing climate.
“This research provides a pathway for making this method cost-effective and scalable,” Peralta-Yahya says. “This system might one day be used to make chemicals ranging from aromatics and terpenes, to alcohols and polymers, and all in a way that not only reduces our carbon footprint, but improves it.”
Funding: Advanced Research Project Agency-Energy (ARPA-E), U.S. Department of Energy and the U.S. Department of Energy, Office of Science, Biological and Environmental Research Program.
News Contact
Written by Selena Langner
Nov. 13, 2024
Facundo Fernández, Vasser Woolley Foundation Chair in Bioanalytical Chemistry and Regents’ Professor in the School of Chemistry and Biochemistry, has been selected as this year’s Anachem Award recipient by the Association of Analytical Chemists.
Presented as a part of a symposium arranged and given by former students and colleagues to honor the recipient, the award recognizes an outstanding analytical chemist that has advanced the field through exemplary research, teaching, or other endeavors.
“This award is very significant to me as it is given to the most accomplished scientists in the field of analytical chemistry, including some of my long-time heroes, such as Bob Kennedy of the University of Michigan, Catherine Fenselau of the University of Maryland and Scott McLuckey of Purdue University,” says Fernández.
“Anachem award winners include Rosalyn Yallow, who received the Nobel Prize for the development of the radioimmunoassay technique,” he adds. “It is enormously significant to be recognized by such close peers who appreciate the value of measurement science in general, and analytical chemistry in particular.”
Fernández is a noted leader in the field of metabolomics and molecular imaging, where his research encompasses the development of new ionization, imaging, machine learning and ion mobility spectrometry tools for probing composition and structure in complex molecular mixtures. He is the author of over 225 peer-reviewed publications and has received the NSF CAREER award, the CETL/BP Teaching award, the Ron A. Hites best paper award from the American Society for Mass Spectrometry, and the Beynon award from Rapid Communications in Mass Spectrometry, among others.
News Contact
Written by Selena Langner
Nov. 12, 2024
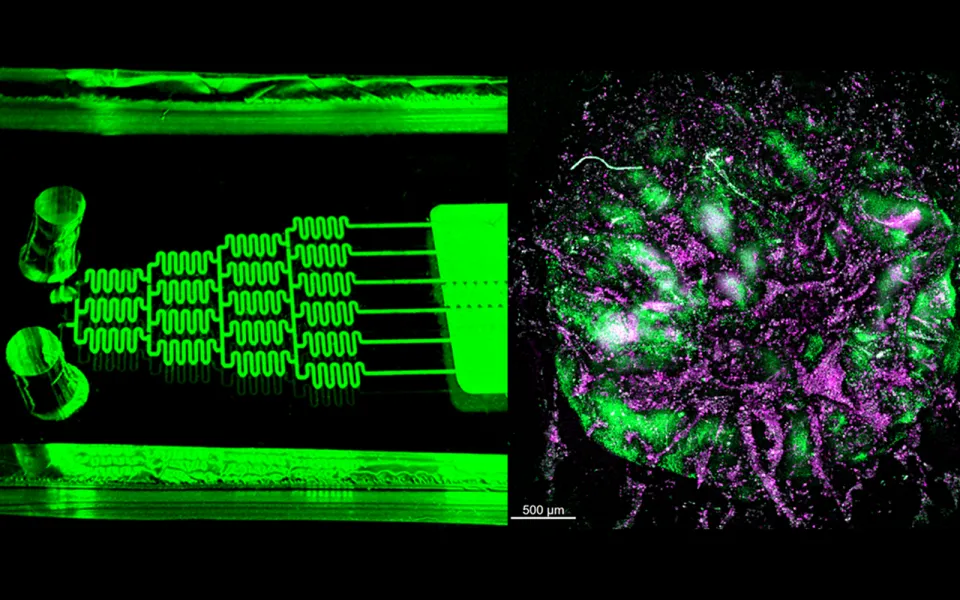
To better understand why some cancer patients struggle to fight off infections, Georgia Tech researchers have created tiny lab-grown models of human immune systems.
These miniature models — known as human immune organoids — mimic the real-life environment where immune cells learn to recognize and attack harmful invaders and respond to vaccines. Not only are these organoids powerful new tools for studying and observing immune function in cancer, their use is likely to accelerate vaccine development, better predict disease treatment response for patients, and even speed up clinical trials.
“Our synthetic hydrogels create a breakthrough environment for human immune organoids, allowing us to model antibody production from scratch, more precisely, and for a longer duration,” said Ankur Singh, Carl Ring Family Professor in the George W. Woodruff School of Mechanical Engineering and professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory.
“For the first time, we can recreate and sustain complex immunological processes in a synthetic gel, using blood, and effectively track B cell responses,” he added. “This is a gamechanger for understanding and treating immune vulnerabilities in patients with lymphoma who have undergone cancer treatment — and hopefully other disorders too.”
Led by Singh, the team created lab-grown immune systems that mimic human tonsils and lymph node tissue to study immune responses more accurately. Their research findings, published in the journal Nature Materials, mark a shift toward in vitro models that more closely represent human immunology. The team also included investigators from Emory University, Children’s Hospital of Atlanta, and Vanderbilt University.
Designing a Tiny Immune System Model
The researchers were inspired to address a critical issue in biomedical science: the poor success rate of translating preclinical findings from animal models into effective clinical outcomes, especially in the context of immunity, infection, and vaccine responses.
“While animal models are valuable for many types of research, they often fail to accurately mirror realistic human immune biology, disease mechanisms, and treatment responses,” said Monica (Zhe) Zhong, a Bioengineering Ph.D. student and the paper’s first author. “To address this, we designed a new model that faithfully replicates the unique complexity of human immune biology across molecular, cellular, tissue, and system levels.”
The team used synthetic hydrogels to recreate a microenvironment where B cells from human blood and tonsils can mature and produce antibodies. When immune cells from healthy donors or lymphoma patients are cultured in these gel-like environments, the organoids support longer cell function, allowing processes like antibody formation and adaptation to occur — similar to the human body. Utilizing the organoids for individual patients helps predict how that individual will respond to infection.
The models also enable researchers to control and test immune responses under various conditions. The team discovered that not all tissue sources are the same, and tonsil cells struggled with longevity issues. They used a specialized setup to study how healthy immune cells react to signals that help them fight infections, which failed to trigger the same response in cells from lymphoma survivors who seemingly have recovered from immunotherapy treatment.
Using organoids embedded in a novel immune organ-on-chip technology, the team observed that immune cells from lymphoma survivors treated with certain immunotherapies do not organize themselves into specific “zones,” the way they normally would in a strong immune response. This lack of organization may help explain some immune challenges cancer survivors face, as evidenced by recent clinical findings.
A Game-Changing Technology
This research is primarily of interest to infectious disease researchers, cancer researchers, immunologists, and healthcare professionals dedicated to improving patient outcomes. By studying these miniature immune systems, they can identify why current treatments may not be effective and explore new strategies to enhance immune defenses.
"Lymphoma patients treated with CD20-targeted therapies often face increased susceptibility to infections that can persist years after completing therapy.Understanding these long-term impacts on antibody responses could be key to improving both safety and quality of life for lymphoma survivors,” said Dr. Jean Koff, associate professor in the department of Hematology and Oncology at Emory University’s Winship Cancer Institute and a co-author on the paper.
“This technology provides deeper biological insights and an innovative way to monitor for recovery of immunological defects over time. It could help clinicians better identify patients who would benefit from specific interventions that reduce infection risk,” Koff added.
Another critical and promising aspect of the research is its scalability: An individual researcher can make hundreds of organoids in a single sitting. The model’s capability to target different populations — both healthy and immunosuppressed patients — vastly increases its usability for vaccine and therapeutic testing.
According to Singh, who directs the Center for Immunoengineering at Georgia Tech, the team is already pushing the research into new dimensions, including developing cellular therapies and an aged immune system model to address aging-related questions.
“At the end of the day, this work most immediately affects cancer patients and survivors, who often struggle with weakened immune responses and may not respond well to standard treatments like vaccines,” Singh explained. “This breakthrough could lead to new ways of boosting immune defenses, ultimately helping vulnerable patients stay healthier and recover more fully.”
The work was initially funded by the Wellcome Leap HOPE program. This support has led to a boost in recent funding, including a recent $7.5M grant from the National Institute of Allergy and Infectious Diseases.
Citation: Zhong, Z., Quiñones-Pérez, M., Dai, Z. et al. Human immune organoids to decode B cell response in healthy donors and patients with lymphoma. Nat. Mater. (2024).
DOI: https://doi.org/10.1038/s41563-024-02037-1
Funding: Wellcome Leap HOPE Program, National Institutes of Health, National Institute of Allergy and Infectious Diseases, National Cancer Institute, and Georgia Tech Foundation
News Contact
Catherine Barzler, Senior Research Writer/Editor
Institute Communications
Oct. 10, 2024
Levi Phillips' path to enrollment as a first-year major in Georgia Tech’s School of Chemical and Biomolecular Engineering (ChBE) in fall 2024 included interning in the Frugal Science Academy, which a ChBE professor created to nurture the next generation of engineers, inventors, and change makers.
Phillips learned of the Frugal Science Academy (FSA) as a student at Lambert High School through his biotechnology teacher Janet Standeven, who had developed a collaboration with FSA founder Saad Bhamla, an associate professor in ChBE.
“The Frugal Science Academy is building a world where synthetic biology is accessible to everyone who wishes to participate through the creation of affordable equipment,” Bhamla said. “We help high school students build synthetic biology tools using everyday objects and share those technologies with others.”
Frugal innovations that have emerged from the Bhamla lab include an automated tracking microscope, a 3D-printed centrifuge, and an inexpensive cell lysis device for molecular biology.
During his senior year at Lambert in Suwanee, Georgia, Phillips worked remotely with Bhamla’s FSA conducting an independent project on a PIO reactor, an affordable open-source bioreactor.
FSA Boot Camp and Teacher Training
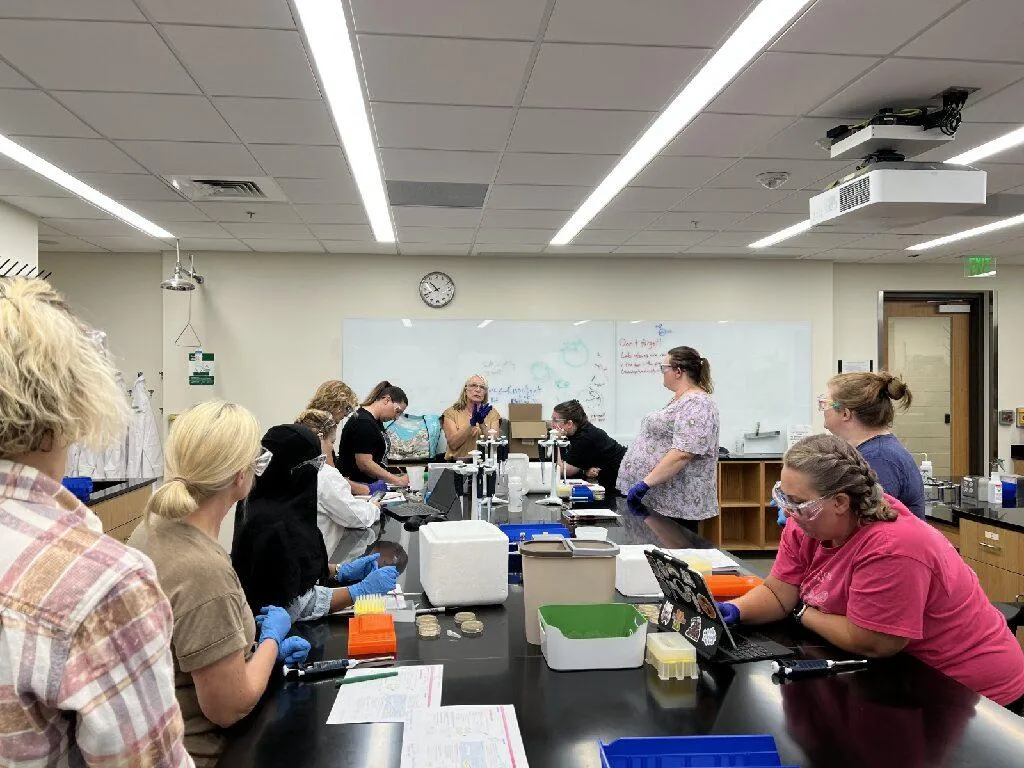
Phillips’ project continued in summer 2024 as he worked in campus labs at Georgia Tech as part of the two-week FSA Boot Camp and Teacher Training, which involved, at various times, 15 high school interns and 20 other high school students working with undergraduate mentors on various projects from new hardware devices to innovative synthetic biology projects.
Six high school teachers also took part in the teacher development portion of the Boot Camp.
Phillips, who said the overall experience with the FSA attracted him to apply and enroll at Georgia Tech, described the mentorship and lab experiences during the Boot Camp as exceptional.
“If I had to give a star rating, it would be six stars out of five, just fantastic,” he said.
Formation and Funding of FSA
Phillips’ former biotechnology teacher at Lambert, Janet Standeven, organized the Boot Camp in her role as program manager of the FSA.
She began partnering with ChBE’s Professor Bhamla in 2017 when her Lambert students wanted to take part in the International Genetically Engineered Machine Competition. Standeven reached out to Bhamla to see if her students could use an open-source centrifuge from his lab for a project.
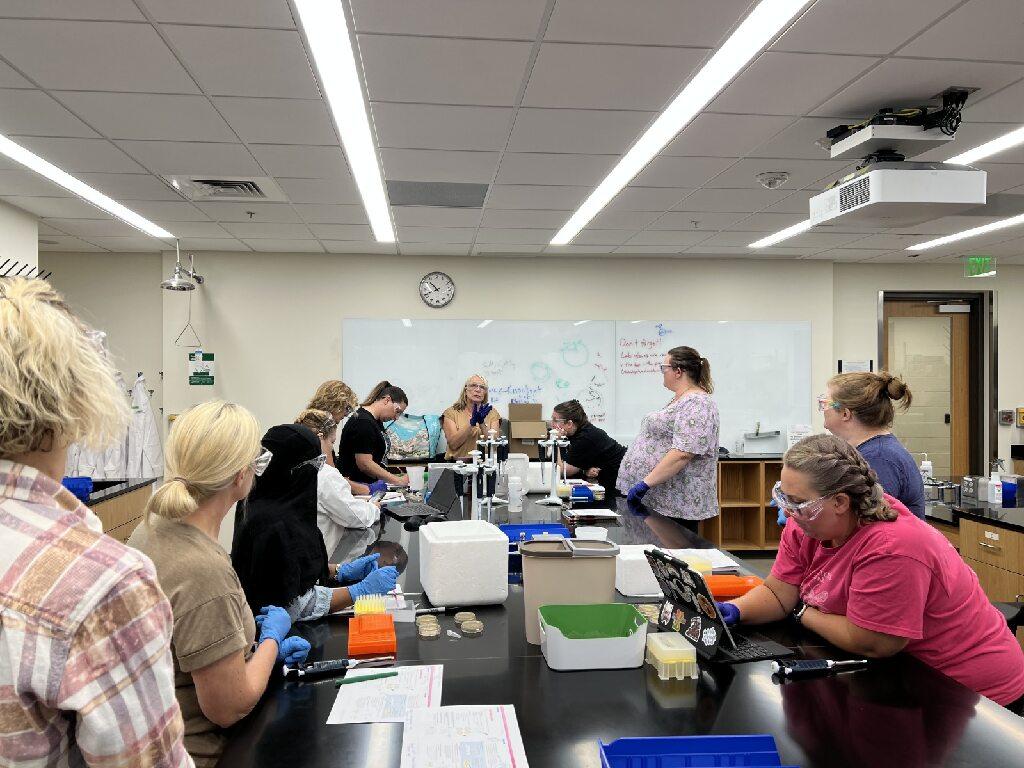

Teachers from across Georgia participated in the Frugal Science Academy Boot Camp and Teacher Training.
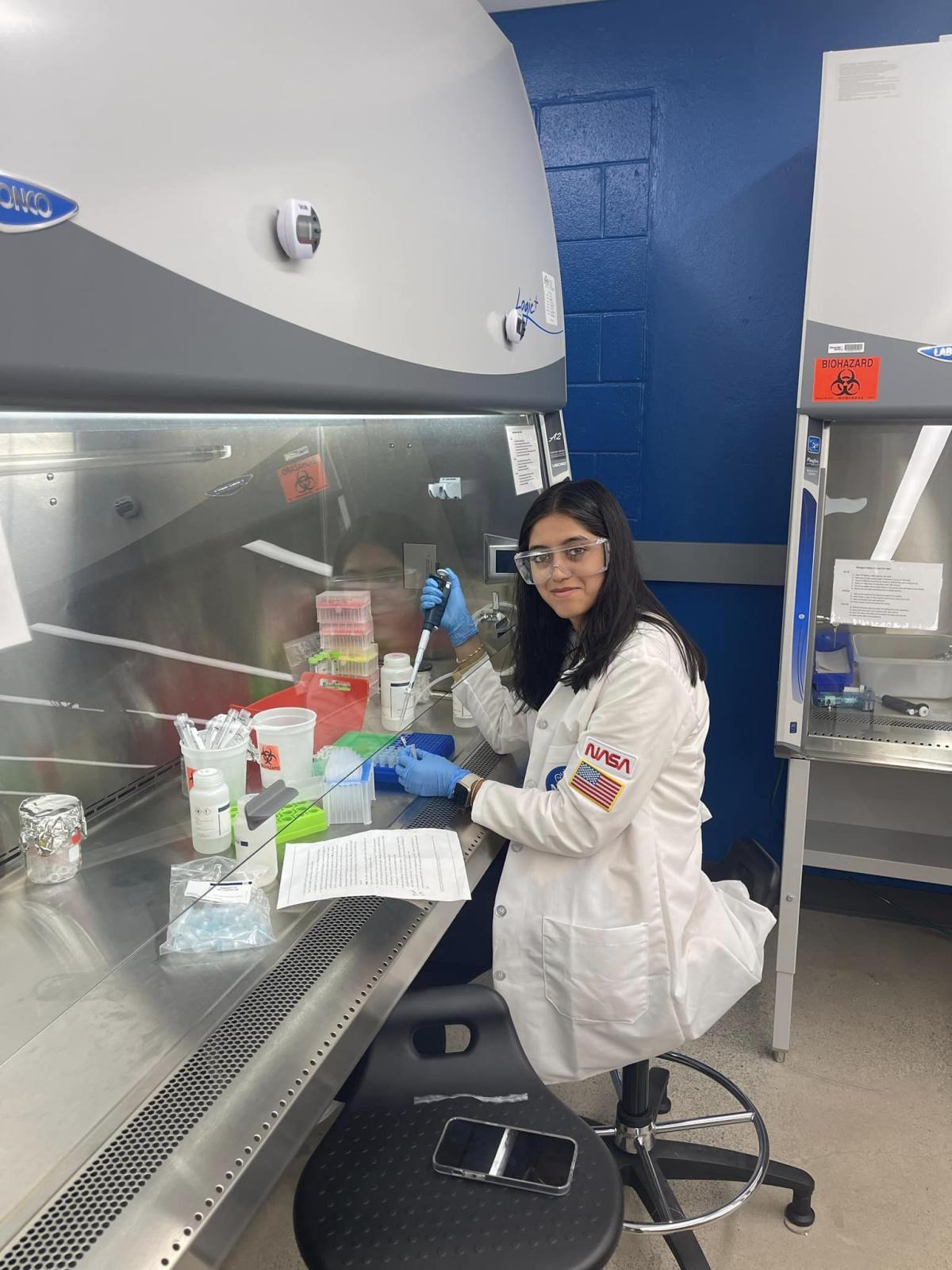

Undegraduate mentors assisted the 35 high school students participating the Summer Boot Camp and Teacher Training
Through the years, their partnership continued and evolved into Standeven, a former Georgia Biotechnology Teacher of the Year, joining the FSA in ChBE full time in 2023. Earlier this year, Georgia Tech's Center for Teaching Learning recognized FSA with the Education Partnership Award.
A five-year Science Education Partnership Award from the National Institutes of Health has enabled Bhamla and Standeven to expand frugal science access beyond Lambert to high schools across Georgia.
In addition to students, the grant focuses on training teachers on how to conduct academic-level research from anywhere using frugal equipment. Teachers in rural areas often do not have access to university research labs that those in Atlanta and other larger cities might.
“We will grow a STEM-engaged workforce by enabling teachers to provide immersive research opportunities at their high schools.” Standeven said. “We’ve found teachers are the multiplier of shared resources. When you train teachers in rapidly evolving technology, they can bring real-life examples into the classroom, enriching their teaching content and student understanding.”
IGEM Competition Preparation
A major focus of the teacher and student training in the summer 2024 FSA Boot Camp included the frugal technologies of a Lambert High School project for the International Genetically Engineering Machine Competition (IGEM).
Lambert is one of only eight high schools in the country to have a team competing in IGEM, which is typically intended for college students.
From 2018 -2023, Lambert IGEM teams won gold medals at the Grand Jamboree International Competition. In 2022, the Lambert team was awarded The Grand Prize for the best high school project in the world for their early diagnostic tool detecting micro RNA associated with Coronary Artery Disease.
Lambert’s current team, consisting of 24 students, will compete at the 2024 international competition in October in Paris, France.
Combating Antimicrobial Resistance
This team is developing a multi-faceted approach (called SHIELD) for combating the growing threats posed by antimicrobial resistance, which can be accelerated by the misuse of antibiotics in agriculture.
One component of SHIELD is ThermoX, a portable device for diagnosing bacterial diseases quickly. Another is a CRISPR-interference (CRISPRi) system designed to target and suppress essential genes in antibiotic-resistant bacteria without using traditional antibiotics.
To measure the effectiveness of CRISPRi in real-world scenarios, the team designed a biosensor called a "toehold," which helps quantify CRISPRi's impact by detecting changes in bacterial genes. Team members also investigated local water contamination caused by antibiotic runoff from livestock, highlighting how this practice fosters resistance.
Lambert senior Christiana Cho, a member of the IGEM team, said that the opportunity to use Georgia Tech’s labs over the summer was a great experience and huge help in advancing their project.
“We didn’t have to cram all of all experiments into the fall, so the boot camp was great in getting our project moving and ensuring that we have results,” she said. “We’re able to get everything set in time for the IGEM competition.”
Valuable Lab Time
Kate Sharer, a biotechnology teacher and IGEM advisor at Lambert, reiterated the value of the lab time made possible by the generosity of the School of Biological Sciences’ lab manager Alison Onstine, who provided space in the Boggs building.
“To have a solid eight hours a day in the lab is something we never get in high school,” said Sharer, a Georgia Tech alum (CHEM 1996).
“To have that has been wonderful as well as the collaborations with the undergraduate mentors and access to the postdocs and faculty members who’ve been generous with their time and expertise.”
Onstine said: “The biology labs are delighted to support these valuable and impactful outreach programs. We are fortunate to have a set of newly renovated teaching labs in the Boggs building that can easily accommodate summer programs and special events. That this program is having such an outsized impact on science education in the local community is all the better.”

High school students in the Frugal Science Academy presented their work in a poster showcase at the end of the two-week Boot Camp.
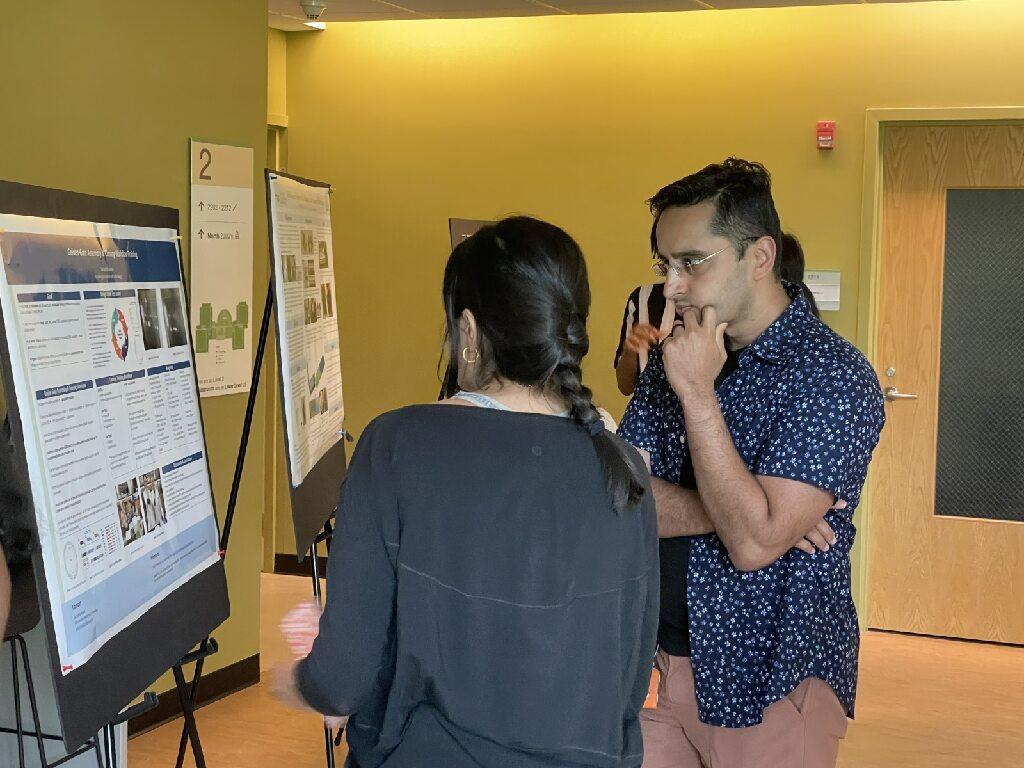

Frugal Science Academy Founder Saad Bhamla learns about a student's research project in the poster showcase.
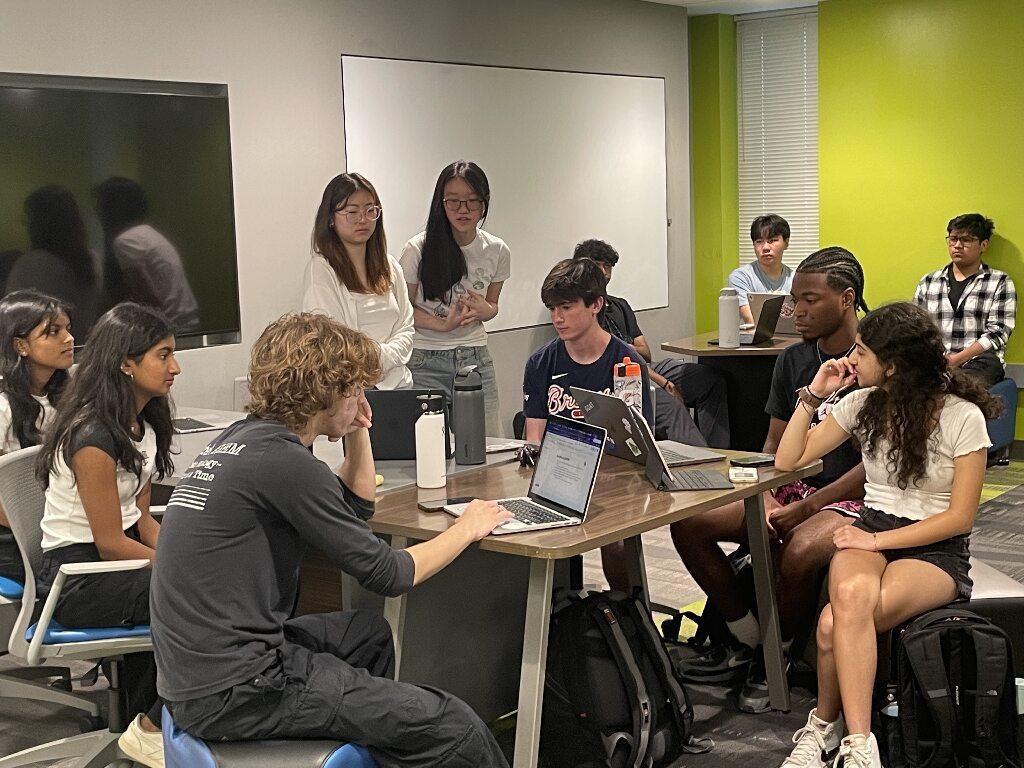

Thirty-five high school students participated in the Boot Camp.
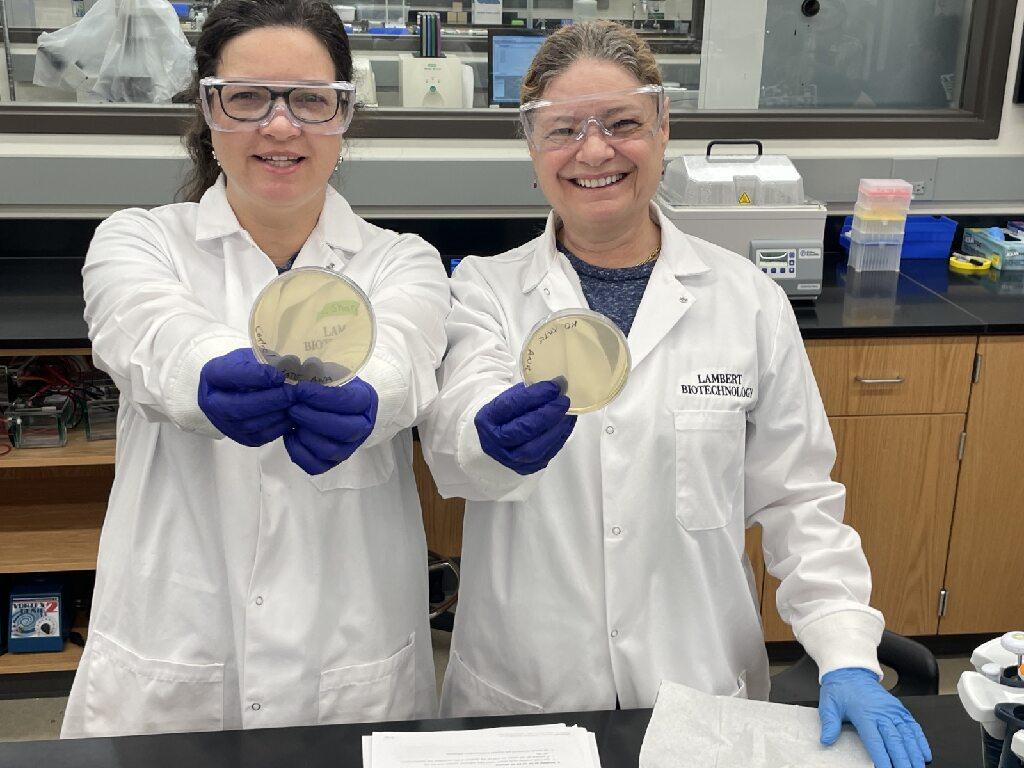

The Frugal Science Academy instructs teachers on how to conduct academic-level research from anywhere using frugal equipment.
Making Impact Statewide
Melissa Rowe, a teacher at Chattooga High School in Summerville, Georgia, attended the FSA boot camp in both 2023 and 2024.
“It’s really increased the rigor of my classes and provided my students with new lab experiences, skills, and knowledge,” she said. “My goal is to help my students understand the topics or concepts of science are not standalone but interconnected.”
She said that learning lab techniques and technology involved in the IGEM student project on preventing antimicrobial resistance (aggravated by the misuse of antibiotics in agriculture) was of particular interest to her because of personal background. She lives on her family’s farm, and many of the people in her community grow some of their own crops.
“Future Farmers of America and 4-H are big in our area,” Rowe said. “I’m planning to apply for grants to test soil from different areas for antibiotic resistance. That’s important to us in the health of our crops and farm animals in our area.”
Rowe said she is grateful to be part of the growing impact that FSA is making on science education across Georgia. “It’s been an amazing experience coming and working in the labs at Georgia Tech.”
Sharing FSA Resources
FSA student projects are added to a website open-sources that other classrooms can access for their experiments. The formation of virtual networks of teachers and classrooms to test new designs and give critiques to the inventors will provide a feedback loop to improve the projects as well as data for possible publications.
“Students at every high school need and deserve these opportunities,” Standeven said. “Eventually, every high school will be able to support a bio-engineering mindset and the opportunity to develop ideas into projects. Eventually, communities will see that synthetic biology is doable, achievable, and a way to solve major world problems.”
News Contact
Brad Dixon
Nov. 12, 2024
Pinar Keskinocak has been selected as the next leader of Georgia Tech’s H. Milton Stewart School of Industrial and Systems Engineering (ISyE). She will serve as the H. Milton and Carolyn J. Stewart School Chair beginning January 1.
Keskinocak is the William W. George Chair and Professor and serves as ISyE’s associate chair for faculty development.
She will be ISyE’s ninth permanent chair, leading a school renowned for its top-ranked graduate and undergraduate industrial engineering programs. U.S. News & World Report has consistently ranked ISyE as the nation's best since the mid-1990s.
“Pinar is a proven and respected leader both on campus and within her academic and research community,” said Raheem Beyah, dean of the College of Engineering and Southern Company Chair. “She is well-positioned to continue advancing ISyE’s national prominence and accelerate the School’s trajectory.”
Keskinocak is the cofounder and director Georgia Tech’s Center for Health and Humanitarian Systems, an interdisciplinary research center focused on education, outreach, and developing innovative solutions via advanced modeling, analytics, and systems engineering.
Keskinocak’s research has had broad societal impact. This includes policies and practices for improved emergency preparedness and response, disease prevention and public health, healthcare access, resource allocation, and supply chain management.
Keskinocak has collaborated with the Centers for Disease Control and Prevention, The Carter Center, and other governmental and nongovernmental organizations to translate research into real-world solutions that benefit people and communities.
“I am honored to have the privilege of serving our School in this important leadership role,” Keskinocak said. “As ISyE continues to expand our core activities in education and research, we will strive to advance our excellence and leadership and grow our impact. I look forward to collaborating with our faculty, staff, students, and alumni, as well as with the leadership of the College, Georgia Tech, and our broader community and partners.”
A highly regarded researcher, Keskinocak has published extensively in top-tier academic journals. She served in various leadership roles within professional societies, including as the 2020 president and a two-time board member of INFORMS (The Institute for Operations Research and Management Sciences). She is the cofounder, and has been president, of multiple INFORMS subdivisions. She also has served on several National Academies of Sciences, Engineering, and Medicine committees.
Keskinocak is a fellow of INFORMS and recipient of the society’s George E. Kimball Medal, President’s Award, and Daniel H. Wagner Prize. At Georgia Tech, she has been recognized with the Outstanding Achievement in Research Program Development Award, Class of 1934 Outstanding Service Award, Outstanding Professional Education Award, and Denning Award for Global Engagement.
A dedicated mentor, educator, and advocate for broadening participation in STEM fields, Keskinocak served as the College’s ADVANCE Professor from 2014 to 2020. She was recognized with the INFORMS Women in OR/MS Award and the Georgia Tech Women in Engineering Excellence Teaching Faculty Award.
Keskinocak replaces Edwin Romeijn, who will return to the ISyE faculty after 10 years as chair.
“I am thankful to Edwin for his very successful tenure, during which ISyE enrollment grew from 1,800 students to more than 8,000,” Beyah said. “I’m also grateful to our search committee and chair Arijit Raychowdhury. This group of students, faculty, and staff diligently worked to help identify a national, diverse pool of strong candidates.”
News Contact
Jason Maderer
Nov. 08, 2024
Georgia Tech School of Electrical and Computer Engineering (ECE) Ph.D. graduate Jesus Antonio Sanchez Perez won the Best Paper Award at the IEEE Engineering in Medicine and Biology Society Body Sensor Networks (BSN) Conference held in Chicago from Oct. 15-17.
Sanchez Perez, who recently started as an assistant professor at his undergraduate alma mater, the University of Puerto Rico at Mayaguez (UPRM), conducted the research in Professor Omer Inan’s Inan Research Lab. It’s the second consecutive year a researcher from Inan’s lab won the Best Paper Award from the conference.
The award-winning research introduced a novel approach for the continuous and non-invasive assessment of pleural effusion (PE) using transthoracic bioimpedance continuously measured at multiple frequencies.
PE, which affects over 1.5 million patients in the U.S., is the accumulation of excess fluid around the lungs, compressing them, increasing the risk of developing infections, and leading to respiratory distress.
Current PE assessments remain limited to sporadic measurements due to their cost, availability, or radiation exposure. This results in the unsatisfactory prognosis and assessment of symptoms for patients.
Sanchez Perez’s approach addresses the problem by exploiting the information embedded in the multi-frequency bioimpedance signals at multiple resolutions, namely the baseline and respiratory components.
He achieved this by extracting six fluid-tracking surrogates and found them to be strongly and significantly correlated to the infused pleural fluid in a porcine animal model of bilateral PE. The early results have promising, paving for continued research and testing.
Ultimately, the research may enable continuous and convenient respiratory monitoring in at-home and uncontrolled clinical settings, which can prevent acute deteriorations, improve treatment outcomes, and help patients.
The technology also has future applications in assessing many diseases including congestive heart failure, liver cirrhosis, and pneumonia.
Sanchez-Perez’s blended his research at ECE with his minor, biomedical engineering. It focused on investigating novel cardiopulmonary monitoring technologies, with a special focus on applications to clinical and acute-care settings.
It helped him to achieve his dream of returning to UPRM as a faculty member, where he’ll now continue doing innovative research with hopes of continuing to help people and achieve better medical outcomes.
News Contact
Zachary Winiecki
Nov. 07, 2024
Programmed Cell Death-1, or PD-1, has become a headline-grabbing molecule best known for its role in cancer immunotherapies called checkpoint inhibitors.
A study from Georgia Tech and Emory University researchers is offering improved understanding of why these inhibitors work — and how to make them effectively fight cancer for more patients.
In a normal, healthy body, PD-1 is a receptor protein that serves as an important off-switch, or checkpoint. Found on a cell’s surface, it binds with a ligand — either PD-L1 or PD-L2 — on another cell surface. This interaction signals the immune systems’ T cells not to attack healthy cells. But sometimes, invading cancer cells also carry a ligand that will bind with PD-1, fooling the body’s immune system into calling off the attack when T cells are needed most.
PD-1 blockade therapy is a checkpoint inhibitor that blocks this signaling process, unleashing the full fury of T cells. Still, only 20% to 40% of patients receive clear benefits from this kind of immunotherapy.
Here’s the thing: Part of what remains unclear is how PD-1 initiates the stand-down order to T cells. Sometimes, ligands bind with PD-1 and don’t suppress T cell activity. So, solving the mystery of what else causes PD-1 to work as a checkpoint can open the door to more effective cancer therapies.
Wallace H. Coulter Department of Biomedical Engineering researcher Cheng Zhu and his collaborators may have found a key: physical force.
“Mechanical forces are an important but previously overlooked component of immunology in general, and specifically in PD-1 activity,” said Zhu, professor Regents' Professor and J. Erskine Love Jr. Chair. “They play a critical role in regulating immune responses.”
Zhu and his team presented their research in Nature Communications, demonstrating that PD-1 is not activated just through interacting or binding with ligands. These ligands must also be anchored to a surface, like a cell membrane, which enables T cells to exert small but measurable forces.
“We show that T cells exert force on this interaction between PD-1 and the ligand. Without that reactive force on the cancer cell ligand, PD-1 doesn’t function,” Zhu said.
And that means T cells can be fully armed and ready for the fight.
Measuring the Force
The recent study is a sequel to research Zhu’s team published in 2021 that explained PD-1’s suppressive role and its value in immunotherapy. This time, the researchers went deeper, identifying and measuring the physical force involved in PD-1’s function.
They used special tools called molecular tension probes and biomembrane force probes to evaluate the tiny physical force exerted by T cells and understand the connection between this molecular jostling and PD-1 activity.
On a solid cell membrane with the appropriate ligand (PD-L1 for example), T cells exert a force of 4.7 to 12 piconewtons. When the force applied was less than 7 piconewtons, the bond between PD-1 and the ligand got stronger and lasted longer. When the force was higher than 8, the bond weakened and broke more easily.
“If the force is too high, the bond breaks, and that weakens PD-1’s ability to stop T cells,” Zhu explained. “It’s a molecular balancing act, and the right amount of physical force makes all the difference.”
While they confirmed that T cells exert small forces on PD-1 attached to a surface-bound ligand, they also discovered that soluble PD-L1 ligands floating freely in the bloodstream lack the mechanical support needed to activate PD-1.
“This explains why soluble ligands don’t trigger T cell inhibition in the same way,” Zhu said.
Ultimately, the research showed that immune cells need physical as well as chemical cues to properly manage PD-1 activity, and even the tiniest show of force could play a role in our body’s ability to protect itself, against cancer and potentially other diseases.
“Next, we would like to test our in vitro findings in an in vivo setting, using animal models,” Zhu said.
CITATION: Kaitao Li, Paul Cardenas-Lizana, Jintian Lyu, Anna V. Kellner, Menglan Li, Peiwen Cong, Valencia E. Watson, Zhou Yuan, Eunseon Ahn, Larissa Doudy, Zhenhai Li, Khalid Salaita, Rafi Ahmed, Cheng Zhu. “Mechanical force regulates ligand binding and function of PD-1.” Nature Communications. doi.org/10.1038/s41467-024-52565-2
FUNDING: This research was supported by the National Science Foundation, grant No. MCA08X014, and the National Institutes of Health, grant Nos. R01CA243486, U01CA250040, U01CA250040S2, RM1GM145394, and F31CA243502. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of any funding agency.
News Contact
Oct. 24, 2024
By Jerry Grillo
Some cancers are stubbornly resistant to the drugs designed to kill them, and Ahmet Coskun wants to know why. So, the Georgia Tech biomedical engineer and his research team has developed a precise molecular tool to help clear up the mystery.
Their graph-based super-resolution protein-protein interaction (GSR-PPI) technique creates a detailed, three-dimensional map of protein interactions within single cells. And it could lead to a better understanding and treatment of drug-resistant cancers.
Coskun will share the results of their work with the rest of the biomedical engineering world this week. As one of this year’s 12 Young Innovators of Cellular and Molecular Bioengineering (CMBE), he’s been invited to present a paper about his research on Friday, Oct. 25, at the 2024 Biomedical Engineering Society (BMES) Annual Meeting in Baltimore.
“Cancer treatments fail in many individuals and pinpointing what goes on at the single cell level is paramount,” said Coskun, assistant professor in the Wallace H. Coulter Department of Biomedical Engineering, where he holds the Bernie Marcus Early Career Professorship.
He noted that while sequencing and molecular imaging have shed light on the molecules associated with disease, there is no definitive test available to decode the communication between two interacting molecules that lead to drug resistance.
“At BMES we’ll showcase a novel direction that creates a high-definition view of lung cancers in individual cells using a graphical representation of molecular wiring diagrams,” he said.
Current methods don’t adequately capture the detailed spatial interactions of proteins in cells, which is crucial for understanding drug resistance. Coskun’s team used single-cell spatial proximity ligation assays and advanced microscopy to examine protein interactions in lung cancer cells treated with the drug Osimertinib. Deep learning models analyzed these images to classify the drug treatment states of the cells.
GSR-PPI significantly outperformed the traditional methods in predicting how cells respond to the drug. It classified how drugs performed in both cancer cells and human lung tissues with high accuracy and differentiated between single and combination drug therapies.
“Basically, it’s like watching a 4k movie of cancer,” said Coskun, the first person from Georgia Tech to receive the Young Innovator honor from CMBE, a leading research journal, in more than a decade.
“Getting acknowledged by peers and mentors is an affirming feeling,” he said. “Our curiosity driven projects, and their translational impacts are being recognized. meets expectations. It’s a sign of my group’s passion to take chances and try new things."
News Contact
By Jerry Grillo
Pagination
- Previous page
- Page 14
- Next page