Dec. 18, 2024
From combating cancer and infections to storing energy, lipid-protein interactions are critical to biological processes in cells. But the mechanisms that drive these interactions have historically been difficult to map and understand.
A study led by Georgia Tech is showcasing a new resource to help researchers understand the structure and function of these interactions — called assemblies — at both molecular and functional levels. The work is published in the Nature-family journal Communications Chemistry.
Called BioDolphin — short for Biological Database of Lipid-Protein Highly Inclusive Interactions — the resource is the first comprehensive, annotated database of protein-lipid interactions. Integrated into a user-friendly web server, BioDolphin is freely accessible to all. Users can easily view and download interaction data and systematically analyze lipid-protein assemblies.
“Understanding lipid-protein interactions is crucial in advancing our understanding of human health and disease treatment,” says the study’s corresponding author, Andrew McShan. “BioDolphin is the first resource to collect this type of information for all kinds of proteins, not just those found in membranes. And because it is publicly available, this information is now at the tips of researchers’ fingertips.”
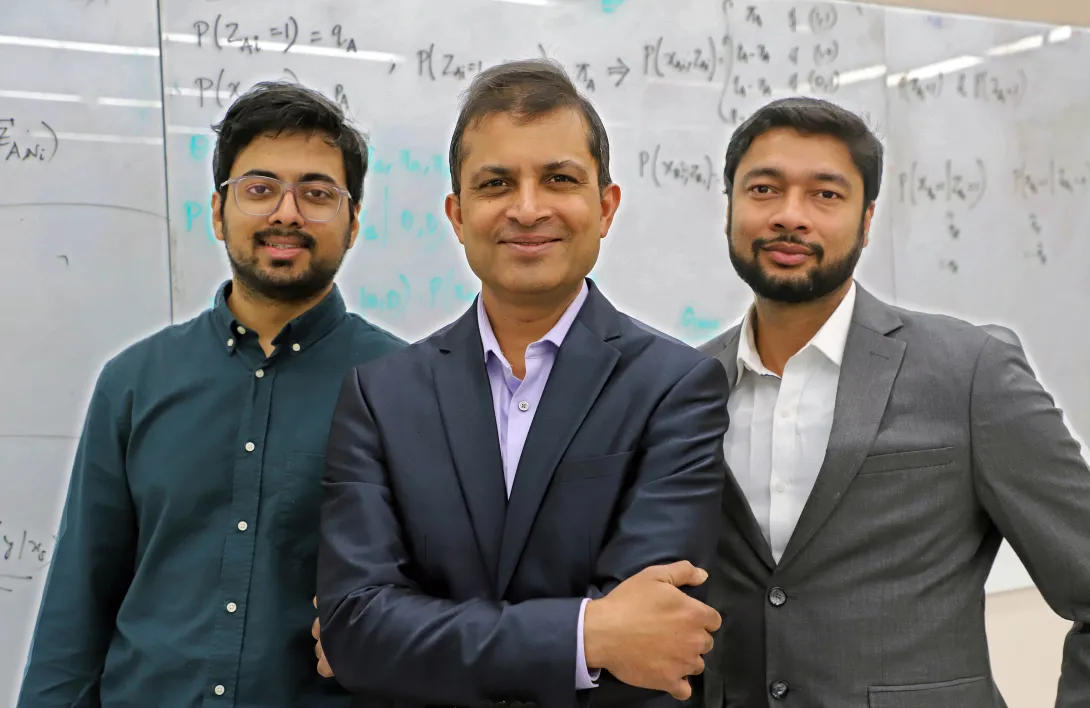
“BioDolphin as a comprehensive database of lipid–protein binding interactions” is led by McShan, an assistant professor in the School of Chemistry and Biochemistry at Georgia Tech, alongside first author Li-Yen (Zoey) Yang, Bioinformatics Ph.D. student; School of Computational Science and Engineering Assistant Professor Yunan Luo; and Kaike Ping, a Ph.D. student at Virginia Tech.
Diving into accessible data
A curated database with richly annotated information, BioDolphin contains over 127,000 lipid-protein binding interactions. And while most databases of lipid-protein assemblies have focused solely on a specific type of protein — membrane proteins — BioDolphin expands beyond that.
“BioDolphin enables us to globally define the structural features of lipid-protein assemblies across the eight different classes of lipid compounds to understand their cellular function and roles in disease,” says McShan, adding that the database also provides information on paired lipid-protein annotation, experimental binding affinities, intermolecular interactions, and atomic structures across a wide range of lipid-protein interactions — all available to anyone with an internet connection.
A molecular blueprint for research — and teaching
“In the past, this research has been limited because lipids are notoriously difficult to study in the lab,” McShan says. "BioDolphin changes the paradigm. It is the first time that anyone has collected, annotated, and analyzed the known structural universe of lipid-protein interactions across all organisms.”
It’s a rapidly developing field. McShan was recently awarded a prestigious Curci grant for cutting-edge cancer research into lipid-based universal immunotherapies and vaccines.
Beyond research applications, the team hopes that BioDolphin will be a resource for biochemistry students.
“The database can serve as a tool for teachers and students studying these protein-lipid interactions, which is often an underdeveloped topic in biology and biochemistry courses,” McShan says. “I hope that BioDolphin is a valuable resource for the researchers of today — and that it can also be a building block for the researchers of tomorrow.”
Funding: Shurl and Kay Curci Foundation, NSF Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support (ACCESS) program, NIH National Institute of General Medical Sciences (NIGMS), Partnership for an Advanced Computing Environment (PACE) at the Georgia Institute of Technology, and Taiwan Ministry of Education Government Scholarship to Study Abroad program.
Dec. 13, 2024
The National Science Foundation has released the results of its annual Higher Education Research and Development Survey (HERD), and Georgia Tech has once again moved up again in the rankings.
The survey of U.S. university research and development expenditures places Georgia Tech as No. 16, up from No. 17 last year, and No. 1 among universities without a medical school. The Institute ranks No. 3 for federally funded research, up from No. 5, and is No. 7 for all externally funded research, up from No. 9.
“Georgia Tech's continued ascension in research rankings is a testament to the exceptional dedication and collaboration of our faculty, staff, and research sponsors,” said Tim Lieuwen, interim executive vice president for Research. “This trajectory, particularly our rise in federally and externally funded research, amplifies the confidence our partners have in Georgia Tech tackle society’s biggest challenges.”
The annual survey compiles R&D expenditure data from U.S. colleges and universities with more than $150,000 in research expenditures during a fiscal year. For fiscal year 2023 (July 1, 2022 – June 30, 2023), that included 914 institutions. Overall, U.S. higher-education R&D spending rose by 11.2%, exceeding $108 billion in fiscal year 2023. This is the largest increase since 2003.
Georgia Tech’s R&D spending of $1.45 billion in fiscal year 2023 reflects an impressive 17.9% increase — $219 million more — from the previous year for the entire research enterprise, which includes the Georgia Tech Research Institute (GTRI). GTRI remains the largest contributor to Georgia Tech’s growth and continues to play a major role in the Institute’s research enterprise and in national defense research.
Additionally, Georgia Tech’s R&D expenditures contributed a monumental $1.45 billion to the state of Georgia’s economy, along with continued growth in commercialization efforts that bring technologies out of the lab and into to the world.
Lieuwen said, “I am proud of these standings and even more excited about the possibilities ahead as we continue to drive innovation that benefits our state, the nation, and the world.”
About Georgia Tech’s Office of the Executive Vice President for Research
The Office of the Executive Vice President for Research (EVPR) directs Georgia Tech’s $1.37 billion (FY 2024) portfolio of research, development, and sponsored activities. This includes leadership of the Georgia Tech Research Institute (GTRI), the Enterprise Innovation Institute, nine interdisciplinary research institutes (IRIs) plus research centers, and related research administrative support units: commercialization, corporate engagement, research development and operations, and research administration. Georgia Tech routinely ranks among the top U.S. universities in volume of research conducted.
News Contact
Shelley Wunder-Smith | shelley.wunder-smith@research.gatech.edu
Director of Research Communications
Dec. 13, 2024
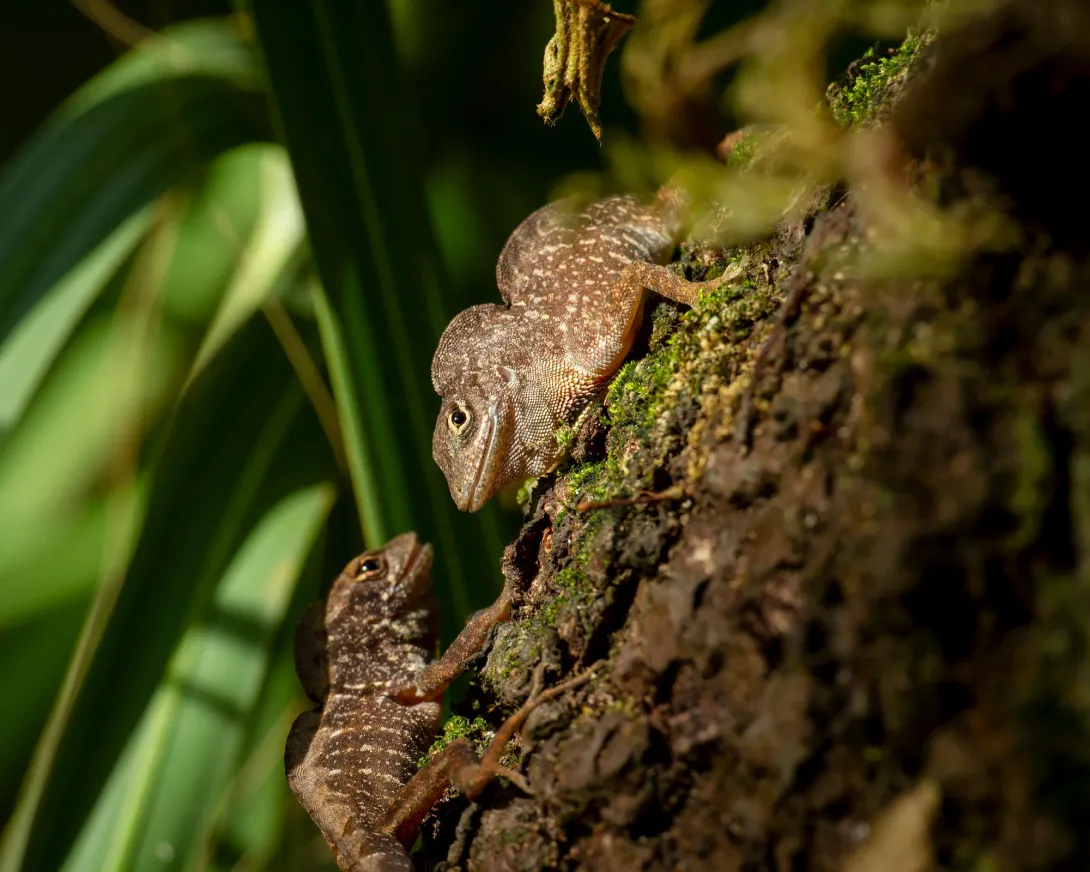
In South Florida, two Caribbean lizard species met for the first time. What followed provided some of the clearest evidence to date of evolution in action.
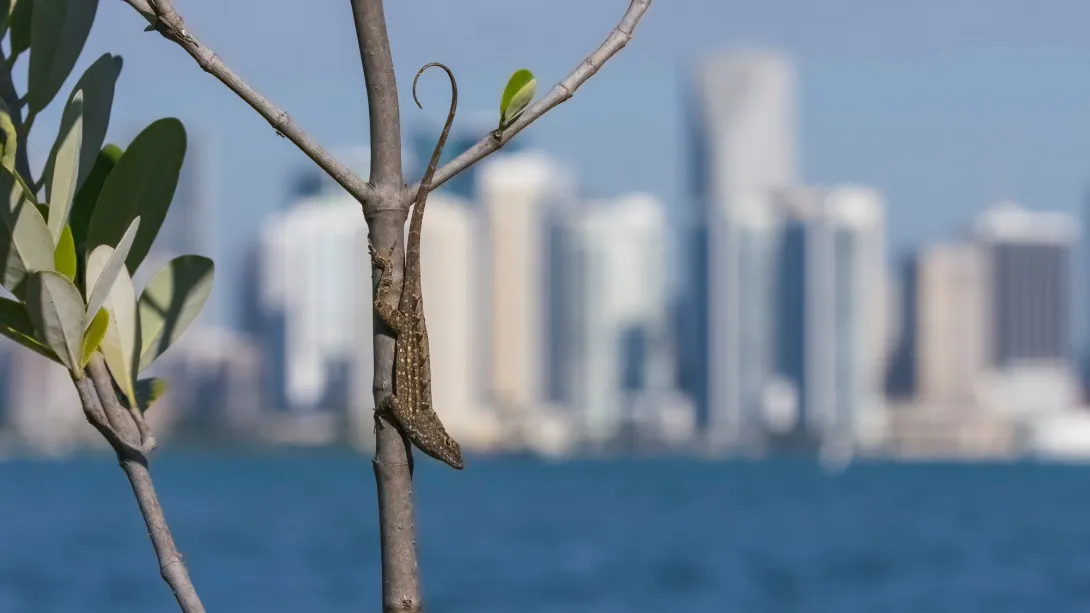
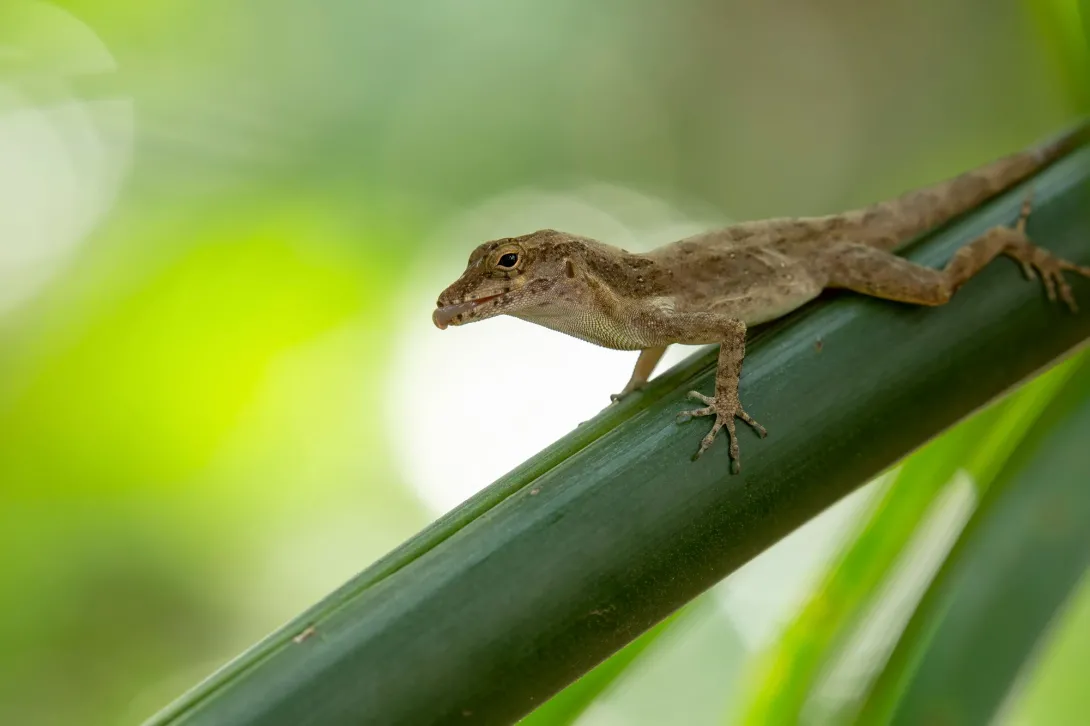
Lead author James Stroud, an assistant professor in the School of Biological Sciences, was studying Cuban brown anoles (Anolis sagrei) in South Florida when the Puerto Rican crested anole (Anolis cristatellus), suddenly appeared in the region.
Published in Nature Communications, the study documents what happens as the two Anolis lizards adapted in response to the new competitor, while helping to resolve a longstanding challenge in evolutionary biology — directly observing the role of natural selection in character displacement: how similar animals adapt in response to competition.
"Most of what we know about how animals change in response to this process comes from studying patterns that evolved long ago,” Stroud says. “This was a rare opportunity where we could watch evolution as it happened."
Competition from coexistence
While these two small, brown lizards diverged evolutionarily between 40-60 million years ago and evolved on completely separate Caribbean islands, the two species are nearly identical, and fill similar ecological niches.
So, when the Puerto Rican crested anole suddenly appeared in Cuban brown anole habitat at Fairchild Tropical Botanic Garden in 2018, the two were competing for similar habitats and food sources.
“When two similar species compete for the same resources, like food and territory, they often evolve differences that allow them to coexist,” Stroud says. But, while scientists have found many examples of similar species developing different traits to ease this overlap, “scientists have rarely been able to observe this process as it unfolds in nature.”
Stroud’s team had already been studying Cuban brown anoles at the Fairchild Tropical Botanic Gardens in Miami, Florida, two years prior to when the crested anoles invaded. The team was able to quickly pivot to observe how the invasion changed both species, analyzing the lizards’ changing diets, measuring if the lizards were moving through foliage or on the forest floor, and recording the different species’ locations relative to each other. For over a thousand lizards, they also measured perch height — the distance from the ground that the lizard is perching — a primary marker of how Anolis lizards divvy up habitat.
“We not only observed how these lizards changed their habitat use and behavior when they encountered each other,” says Stroud, “but we also documented the natural selection pressures driving their physical evolution in real-time."
Human-made habitats and natural experiments
The research team found that when these lizard species occur together, they divide up their habitat in predictable ways — the Cuban brown anole shifted to spend more time on the ground, and evolved longer legs to run faster in this habitat, while the slightly larger Cuban crested anole lived in vegetation above the ground.
"We found that brown anoles with longer legs had higher survival after crested anoles showed up," says Stroud. "This matches perfectly with the physical differences we see in populations where these species have been living together for many generations."
Stroud adds that while the research provides some of the strongest observations of evolution in action to date, it also demonstrates how human activities can create natural experiments that help us understand fundamental evolutionary processes — both species of Anolis lizard in the study were originally non-native to South Florida.
“As species increasingly come into contact due to human-mediated introductions and climate change, these studies may be important for predicting how communities will respond,” he says. "By studying these non-native lizards who are meeting each other for the first time in their existence, we had a unique opportunity to see the actual process unfold and connect it to the patterns we observe in nature."
Dec. 10, 2024
Saurabh Sinha and a multi-institutional team of researchers have created a computational toolkit with the detection power and precision of a spy satellite. But instead of keeping tabs of human traffic on the ground, or infrastructure development in a city, they’re focusing on RNA with unprecedented clarity at the subcellular level.
Their intracellular spatial transcriptomic analysis toolkit, or InSTAnT, can analyze cellular data and chart RNA interactions, providing new insights into the molecular processes of life and advancing an evolving field of research.
“Conventional spatial transcriptomics maps RNA at the tissue level,” said Sinha, professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. “But InSTAnT represents a step forward. It provides, for the first time, an analytic technique to fully exploit single-molecule resolution. This means we can explore the intricate architecture, machinery, and activity of cells in ways that were not possible before.”
In addition to Georgia Tech and Emory, the team included researchers from from the University of Illinois Urbana-Champaign. With Anurendra Kumar, a grad student in the Sinha lab, as lead author, they explained their innovative work recently in Nature Communications.
Subcellular GPS
Spatial transcriptomics has enhanced the study of gene expression (how genes regulate cellular functions and behaviors), revealing molecular activity in its natural environment. The aim is to gain a deeper understanding of biology, health, and disease, with the hope of developing targeted treatments.
“One of the biggest challenges in the field was the lack of systematic tools to analyze spatial relationships at the subcellular level,” Sinha said. “We saw this gap as an opportunity to innovate and solve a problem that was truly spatial in nature.”
InSTAnT was designed to work in tandem with imaging-based spatial transcriptomics technologies like MERFISH (Multiplexed Error-Robust Fluorescence In Situ Hybridization, developed by Harvard in 2015), which can observe thousands of RNA molecules inside single cells, gathering detailed information about gene activity.
“It’s like a GPS for tissue, looking all the way down to city street level,” said Sinha. “The little dots on this GPS aren’t people. They’re RNA molecules called gene transcripts. But we didn’t really know how to make sense of this distribution of molecules in the cytoplasm or the nucleus, or generally within the cell.”
InSTAnT translates what MERFISH gathers, using advanced statistical tests and algorithms, analyzing the distribution of RNA molecules that carry genetic information needed for various cell functions.
The Cities in Our Cells
If a cell was a busy little city, think of the gene transcripts — RNA molecules, the dots in Sinha’s GPS scenario — as workers moving around town, performing their important tasks.
InSTAnT keeps tabs on this activity, investigating where and how these workers interact, and what they might be up to. So, InSTAnT identifies RNA pairs in specific areas, observing molecular interactions that are critical for cellular functions like protein production.
“Our toolkit provides a level of detail crucial for understanding complex biological processes and how they contribute to diseases,” said Sinha, whose team tested the toolkit on a variety of datasets, including human and mouse cells, and across multiple cell types and brain regions.
He expects InSTAnT to transform how researchers study RNA interactions and explore unknown aspects of cellular organization and function.
“I think we’ve opened new possibilities for studying how cells coordinate their activities and adapt to challenges,” said Sinha, adding, “and it was a true team effort, with two other PIs from another institution, and a talented Ph.D. student as the lead author. This is a great example of how collaboration and data-driven science can uncover new biological frontiers.”
CITATION: Aunrendra Kumar, Alex Schrader, Bhavay Aggarwal, Ali Ebrahimpour Boroojeny, Marisa Asadian, JuYeon Lee, You Jin Song, Sihai Dave Zhao, Hee-Sun Han, Saurabh Sinha. “Intracellular spatial transcriptomic analysis toolkit (InSTAnT),” Nature Communications. https://doi.org/10.1038/s41467-024-49457-w
FUNDING: This research was supported by the National Institutes of Health, grant Nos. R35GM131819, R35GM147420, R21HG013180, and T32- 842 GM136629; Johnson & Johnson (WiSTEM2D Award for Science). Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of any funding agency.
News Contact
Dec. 10, 2024
Kristine Lacek vividly remembers watching news coverage of the West Africa Ebola outbreak while she was in high school. Inspired by the brave scientists investigating the disease, she wrote one of her Georgia Tech application essays about her aspiration to work at the Centers for Disease Control and Prevention (CDC) and contribute to their mission of preventing, detecting, and responding to disease threats.
Less than 10 years later, her high school dream became a hectic, fast-paced — and fulfilling — reality. Armed with an accelerated bachelor’s degree in biology and a master’s degree in bioinformatics from Georgia Tech, the double Jacket started a fellowship at the CDC during a pivotal time in history — the early days of the COVID-19 pandemic.
“It was sink or swim for sure,” says Lacek. “Knowing I was working on public health decision-making that could make a lifesaving difference worldwide showed me I had chosen the right path.”
Today, Lacek continues her drive to make a positive global impact as a bioinformatics scientist at the CDC, specializing in influenza and SARS-CoV-2 genomics. Her career has taken her around the world, with time spent in places like Ghana, Oman, Panama, Algeria, India, Thailand, and the Republic of Georgia. She currently lives in Denver, but will return to Georgia Tech to provide the graduation speech at the College of Sciences’ inaugural master’s commencement ceremony this December.
We recently sat down with Lacek to talk about her career and Georgia Tech experience:
What is your favorite memory from Georgia Tech?
Lacek: I always enjoyed the Georgia Tech nights at various Atlanta special event locations like the aquarium and Six Flags. When I was in grad school, the grad gala was held at the Fernbank Museum of Natural History. Halfway through the night, my then-boyfriend-and-now husband looked around and said we should get married here — and we did!
What were some of your college activities?
Lacek: I worked a lot to fund my way through school. I tutored at the Center for Academic Success and worked as an athletic training aid with the track and field team. I gained research experience in the Exercise Physiology Laboratory as an undergrad and in the Gibson Lab as a grad student. Each summer, I served as the teaching assistant for the Biomolecular Engineering, Science, and Technology (BEST) Study Abroad Program in Lyon, France.
How did Georgia Tech prepare you for success?
Lacek: The mix of coursework in my undergraduate and graduate studies was ideal for my career. As an undergrad biology major, I learned key theories and scientific concepts that I still use daily. Studying bioinformatics in grad school, I refined my technical skills in coding, math, and computer science. My two skill sets work well together. Because I understand the molecular side of the interdisciplinary coin, I can better apply technical tools to get the answers I need from the data.
What advice do you have for Georgia Tech students, particularly those looking for a career in bioinformatics?
Lacek: Being a fast learner is the best skill you can have, especially as technology continues to rapidly evolve. The things you are learning right now may not be the exact language or application you will use as a young professional, so the ability to learn new products, programs, and schema quickly will make you very valuable.
On the public health side of things, I think being a really good collaborator and communicator is quintessential for success. One of my biggest regrets is not learning another language. As someone who does a large amount of technical support for other countries and overseas partners, working well with others and good communication is vital.
How do you define bioinformatics?
Lacek: To me, bioinformatics is like a triangle of biology, computer science, and mathematics/statistics. I’m kind of halfway between the biology and computer science side, focusing a lot on next generation sequencing. I use code and statistical applications to make global health predictions based on the data analytics available.
Tell us more about what you do.
Lacek: I do a lot of genomic surveillance, which is basically tracking and monitoring genetic material to detect new mutations and variants. Influenza, for example, circulates year-round worldwide, and we are constantly sequencing samples from all over the place to track what the virus currently looks like and project what will happen globally. At the same time, we're also monitoring for novel outbreaks, with a posture of pandemic preparedness so that if something new and scary pops up, we are already looking for it.
How are you making a difference in the world today?
Lacek: After the COVID-19 emergency response wound down, I moved my focus to influenza. Over the last two years I have been going around the world to train other ministries of health in bioinformatics and next generation sequencing to do what we do in the United States for respiratory virus surveillance.
I believe I've trained and supported scientists from 89 different countries. Because of this effort, we’ve detected some novel variants, such as a new swine flu in Vietnam. It’s thrilling to know that we are making a worldwide impact by helping countries who don’t always have the technical resources and informatics personnel we enjoy here in the U.S.
What are your hobbies?
Lacek: I love to read; I read 106 books last year! I live in Denver, so of course, I enjoy hiking. I recently completed my first 14er (hiking a mountain peak that’s 14,000 feet above sea level). I also love to thrift, cook, and eat out!
News Contact
Writer: Laura S. Smith
Communications Officer II
College of Sciences
laura.smith@cos.gatech.edu
Dec. 10, 2024
A firm handshake between proteins on immune cells is important for the body’s ability to fight infection. Meanwhile, a weak grip helps explain the poor immune deficiencies caused by a rare genetic disorder.
A new study led by Georgia Tech and Emory University researcher Cheng Zhu explores the role of physical force on the immune system’s ability to fight an infection. The team’s discoveries could lead to new therapies that boost immune responses and improve the outcomes of patients battling a rare and devastating disease.
“With this research, we’ve shown how dynamic and physical the immune system truly is,” said Zhu, Regents' Professor and J. Erskine Love Jr. Chair in the Wallace H. Coulter Department of Biomedical Engineering (BME).
The work focuses on the interaction of B cells and T cells in the body’s immune system via two proteins — CD40 on B cells and CD40L on T cells — in an immune deficiency disease called X-linked Hyper IgM syndrome, or X-HIgM. It’s a genetic disorder affecting two out of every million newborn boys, 80% of whom die before the age of 25.
The researchers found mechanical forces generated by these interactions create a “catch bond” between the proteins. It’s like a strong handshake that only gets firmer when each person tries to pull away.
When the bond is strong, it causes T cells to signal B cells they need to make antibodies to fight an invading pathogen. In fact, the B cells can actually switch gears, producing antibodies that are different from what they had been making.
But people with X-HIgM have damaged CD40L proteins, resulting in weak bonds, poor signaling, and the inability to make the right antibodies.
The team published their findings in Science Advances. The work emphasizes the role of mechanotransduction — the conversion of physical force into chemical activity — in the immune system.
Zhu’s fellow principal investigators in the study included Georgia Tech researcher Ankur Singh and Juergen Wienands of the University Medical Center Göttingen in Germany. Lead authors were BME PhD student Stefano Travaglino and former postdoc Hyun-Kyu Choi (now an assistant professor at Yonsei University in South Korea).
Training Camp for B Cells
In the body’s defense system, B cells are produced in the bone marrow and migrate to a part of the lymph nodes called the germinal center.
“It’s like a training camp where B cells undergo improvement processes, including affinity maturation and antibody class switch, enhancing their ability to make effective antibodies,” Travaglino said.
B cells interact with and receive instructive signals from T cells to make antibodies that are most effective in coping with the pathogen invader. It’s a process that relies heavily on the interaction of CD40 and CD40L.
Using techniques like fluorescence microscopy, the researchers were able to look closely at activity in germinal centers. They used force spectroscopy tools like the biomembrane force probe which revealed that the strong, tugging handshake is suppressed by X-HIgM mutation.
The findings suggest that the physical environment and activity within the germinal center is just as important as the chemical signals at play between the proteins. By demonstrating how X-HIgM mutations impair catch bonds, the researchers provided a mechanistic explanation for the condition’s antibody deficiencies — knowledge that could open the door to future innovations in therapeutic intervention and immunotherapy.
Singh called the team’s findings “nothing short of revolutionary.”
“The significance of the research extends far beyond understanding X-HIgM, offering a fresh perspective on how to approach a variety of immune disorders,” he said. “As this field of study evolves, the potential for advancements in immune therapies looks bright.”
CITATION: Hyun-Kyu Choi, Stefano Travaglino, Matthias Münchhalfen, Richard Görg, Zhe Zhong, Jintian Lyu, David M. Reyes-Aguilar, Jürgen Wienands, Ankur Singh, and Cheng Zhu. “Mechanotransduction governs CD40 function and underlies X-linked Hyper IgM syndrome,” Science Advances. DOI: 10.1126/sciadv.adl5815
FUNDING: This research was supported by National Institutes of Health grants U01CA250040, U01CA280984, R01CA238745, and R01CA266052; The Hyper IgM Foundation AWD-004331; German Research Foundation SFB TRR 274, project A08; National Research Foundation of Korea (NRF) grant RS-2024-00337196; and the Yonsei University Research Fund 2024-22-0036. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of any funding agency.
News Contact
Dec. 03, 2024
A new machine learning (ML) model from Georgia Tech could protect communities from diseases, better manage electricity consumption in cities, and promote business growth, all at the same time.
Researchers from the School of Computational Science and Engineering (CSE) created the Large Pre-Trained Time-Series Model (LPTM) framework. LPTM is a single foundational model that completes forecasting tasks across a broad range of domains.
Along with performing as well or better than models purpose-built for their applications, LPTM requires 40% less data and 50% less training time than current baselines. In some cases, LPTM can be deployed without any training data.
The key to LPTM is that it is pre-trained on datasets from different industries like healthcare, transportation, and energy. The Georgia Tech group created an adaptive segmentation module to make effective use of these vastly different datasets.
The Georgia Tech researchers will present LPTM in Vancouver, British Columbia, Canada, at the 2024 Conference on Neural Information Processing Systems (NeurIPS 2024). NeurIPS is one of the world’s most prestigious conferences on artificial intelligence (AI) and ML research.
“The foundational model paradigm started with text and image, but people haven’t explored time-series tasks yet because those were considered too diverse across domains,” said B. Aditya Prakash, one of LPTM’s developers.
“Our work is a pioneer in this new area of exploration where only few attempts have been made so far.”
[MICROSITE: Georgia Tech at NeurIPS 2024]
Foundational models are trained with data from different fields, making them powerful tools when assigned tasks. Foundational models drive GPT, DALL-E, and other popular generative AI platforms used today. LPTM is different though because it is geared toward time-series, not text and image generation.
The Georgia Tech researchers trained LPTM on data ranging from epidemics, macroeconomics, power consumption, traffic and transportation, stock markets, and human motion and behavioral datasets.
After training, the group pitted LPTM against 17 other models to make forecasts as close to nine real-case benchmarks. LPTM performed the best on five datasets and placed second on the other four.
The nine benchmarks contained data from real-world collections. These included the spread of influenza in the U.S. and Japan, electricity, traffic, and taxi demand in New York, and financial markets.
The competitor models were purpose-built for their fields. While each model performed well on one or two benchmarks closest to its designed purpose, the models ranked in the middle or bottom on others.
In another experiment, the Georgia Tech group tested LPTM against seven baseline models on the same nine benchmarks in zero-shot forecasting tasks. Zero-shot means the model is used out of the box and not given any specific guidance during training. LPTM outperformed every model across all benchmarks in this trial.
LPTM performed consistently as a top-runner on all nine benchmarks, demonstrating the model’s potential to achieve superior forecasting results across multiple applications with less and resources.
“Our model also goes beyond forecasting and helps accomplish other tasks,” said Prakash, an associate professor in the School of CSE.
“Classification is a useful time-series task that allows us to understand the nature of the time-series and label whether that time-series is something we understand or is new.”
One reason traditional models are custom-built to their purpose is that fields differ in reporting frequency and trends.
For example, epidemic data is often reported weekly and goes through seasonal peaks with occasional outbreaks. Economic data is captured quarterly and typically remains consistent and monotone over time.
LPTM’s adaptive segmentation module allows it to overcome these timing differences across datasets. When LPTM receives a dataset, the module breaks data into segments of different sizes. Then, it scores all possible ways to segment data and chooses the easiest segment from which to learn useful patterns.
LPTM’s performance, enhanced through the innovation of adaptive segmentation, earned the model acceptance to NeurIPS 2024 for presentation. NeurIPS is one of three primary international conferences on high-impact research in AI and ML. NeurIPS 2024 occurs Dec. 10-15.
Ph.D. student Harshavardhan Kamarthi partnered with Prakash, his advisor, on LPTM. The duo are among the 162 Georgia Tech researchers presenting over 80 papers at the conference.
Prakash is one of 46 Georgia Tech faculty with research accepted at NeurIPS 2024. Nine School of CSE faculty members, nearly one-third of the body, are authors or co-authors of 17 papers accepted at the conference.
Along with sharing their research at NeurIPS 2024, Prakash and Kamarthi released an open-source library of foundational time-series modules that data scientists can use in their applications.
“Given the interest in AI from all walks of life, including business, social, and research and development sectors, a lot of work has been done and thousands of strong papers are submitted to the main AI conferences,” Prakash said.
“Acceptance of our paper speaks to the quality of the work and its potential to advance foundational methodology, and we hope to share that with a larger audience.”
News Contact
Bryant Wine, Communications Officer
bryant.wine@cc.gatech.edu
Nov. 21, 2024
A multi-institutional team of researchers, led by Georgia Tech’s Francesca Storici, has discovered a previously unknown role for RNA. Their insights could lead to improved treatments for diseases like cancer and neurodegenerative disorders while changing our understanding of genetic health and evolution.
RNA molecules are best known as protein production messengers. They carry genetic instructions from DNA to ribosomes — the factories inside cells that turn amino acids into the proteins necessary for many cell functions. But Storici’s team found that RNA can also help cells repair a severe form of DNA damage called a double-strand break, or DSB.
A DSB means both strands of the DNA helix have been severed. Cells have the tools to make some repairs, but a DSB is significant damage — and if not properly fixed can lead to mutations, cell death, or cancer. (Interestingly, cancer treatments, like chemotherapy and radiation, can cause DSBs.)
Storici, a professor in the School of Biological Sciences, has dedicated her research to studying the molecules and mechanisms underlying damaged DNA repairs. Ten years ago, she and collaborators discovered that RNA could serve as a template for DSB repair.
“Now we’ve learned that RNA can directly promote DSB repair mechanisms,” said Storici, whose lab teamed with mathematics experts in the lab of Nataša Jonoska from the University of South Florida. They’re all part of the Southeast Center for Mathematics and Biology based at Georgia Tech. They explain their discovery in the journal Nature Communications.
“These findings open up a new understanding of RNA's potential role in maintaining genome integrity and driving evolutionary changes,” added Storici.
The researchers used variation-distance graphs to visualize millions of DSB repair events, offering a comprehensive snapshot of sequence variations. The graphs highlighted major differences in repair patterns, depending on the DSB position.
This mathematical approach also uncovered significant differences in repair efficiency, pointing to RNA's potential in modulating DSB repair outcomes.
“These findings underscore the critical role of mathematical visualization in understanding complex biological mechanisms and could pave the way for targeted interventions in genome stability and therapeutic research,” said Jonoska.
Molecular Grunt Work
When a DSB happens in DNA, it’s like a load-bearing beam in a building breaking. A careful, precise repair is needed to ensure the building’s — or the DNA’s — stability. The pieces must be rejoined accurately to prevent further damage or mutation. Repairing a damaged building requires having a reliable foreman on the job site. A DSB requires something very similar.
“A key mechanism we identified is that RNA can help position and hold the broken DNA ends in place, facilitating the repair process,” explained Storici, whose team conducted the research in both human and yeast cells.
Specifically, they found that RNA molecules and the broken section of DNA can match up like puzzle pieces. When RNA has this kind of complementarity with the DNA break site, it acts as a scaffold, or a guide, beyond its traditional coding function, showing the cellular machinery where to make repairs. Over millennia, cells have evolved complex mechanisms to fix DSB, each of them functioning like different tools from the same toolbox.
Storici’s team showed that RNA can influence which tools are used, depending on its complementarity to the broken DNA strands. This means that in addition to being the important protein production messenger, RNA acts as both a foreman and laborer when it comes to DNA repair.
A deeper understanding of RNA’s role in DNA repair could lead to new strategies for strengthening repair mechanisms in healthy cells, potentially reducing the harmful effects of treatments like chemotherapy and radiation.
“RNA has a much broader function than we knew,” Storici said. “We still have a lot of research to do into these mechanisms, but this work opens up new ways for exploring how RNA could be harnessed in healthcare, potentially leading to new treatments for cancer and other genetic diseases.”
As Storici and other researchers continue probing RNA’s effects in DNA repair, their revelations could have a lasting impact on human health and evolution. That means better gene therapies, new cancer treatments and anti-aging strategies — and also the ability to influence how organisms adapt and evolve.
CITATION: Youngkyu Jeon, Yilin Lu, Margherita Maria Ferrari, Tejasvi Channagiri, Penghao Xu, Chance Meers, Yiqi Zhang, Sathya Balachander, Vivian S. Park, Stefania Marsili, Zachary F. Pursell, Nataša Jonoska, Francesca Storici. “RNA-mediated double-strand break repair by end-joining mechanisms.” Nature Communications https://doi.org/10.1038/s41467-024-51457-9
FUNDING: NIH grants GM115927, ES028271; NSF grant MCB-1615335; Howard Hughes Medical Institute Faculty Scholar grant 55108574; Southeast Center for Mathematics and Biology NSF DMS-1764406; Simons Foundation grant 59459; NSF grants CCF-2107267 and DMS-2054321.
News Contact
Nov. 21, 2024
A multi-institutional research initiative aims to address lymphoma survival disparities in African American and EBV-infected patients.
A new interdisciplinary initiative with researchers at Georgia Tech, Emory University, MD Anderson Cancer Center, and Weill Cornell Medical aims to address the knowledge gap in lymphomas — particularly diffuse large B-cell lymphoma (DLBCL), the most common form of blood cancer. Survival rates for DLBCL are lower among African American patients and those with Epstein-Barr virus (EBV), which is prevalent in Latin America. The team uses immunoengineering tools to facilitate this discovery.
Tackling Health Disparities in Lymphoma Treatment
To address these health disparities, the team combines expertise in cancer biology and immunoengineering. At Georgia Tech, Ankur Singh works with oncologists and cancer biologists from partner institutions to create innovative cancer technologies, such as lab-grown, lymph node-mimicking models of DLBDL tumors. Singh is Carl Ring Family Professor in the George W. Woodruff School of Mechanical Engineering and the Wallace H. Coulter Department of Biomedical Engineering (BME) and directs the Center for Immunoengineering. These models will mimic the tumor environments in lymphoma from African American patients and model specific mutations prevalent in these patients. Researchers will observe how various genetic changes work in concert with the immune system to impact a tumor's response to treatments.
“We want to understand the full makeup of these tumors; not just the cancer cells but the surrounding supportive cells and proteins,” said Singh, who serves as co-investigator for LLS SCOR. “This study will help us pinpoint which parts of the tumor are critical for its survival and how we can disrupt those mechanisms, including the immune cells.”
Challenges for Understanding Tumor Biology in High-Risk Groups
Diffuse large B-cell lymphoma is the most common form of blood cancer. While many patients respond well to standard therapies, a significant portion — including a disproportionate number of African Americans and individuals with EBV-related conditions, experience poorer outcomes. The reasons behind these disparities are still largely unknown. Current barriers include a lack of diverse representation in research studies and a paucity of engineered technologies dedicated to understanding cancers in patients from underrepresented backgrounds.
"Most lymphoma studies don't include nearly enough African American or Hispanic patients," said Jean Koff, lead investigator and associate professor of Hematology and Medical Oncology at Emory University’s Winship Cancer Institute. “This means we are likely missing key insights into the unique biology and treatment needs of these populations.”
A Collaboration Focused on Advancing Lymphoma Research and Care
This new initiative, funded by The Leukemia & Lymphoma Society's Specialized Center of Research (SCOR) Program, will analyze a comprehensive collection of DLBCL tumor samples that includes many cases from Black and Hispanic patients. By examining genetic differences and tumor structures, the researchers hope to identify the factors most important for improving therapy for these groups.
“This program is groundbreaking because it addresses both biological and structural barriers in treatment, leveraging the latest bioengineered technologies,” Singh noted. “We’re looking at factors that have been overlooked for too long in cancer research, especially in high-risk communities.”
To explore the composition and diversity of cells within tumors of African American patients and better understand how they grow and respond to treatments, the team leverages the expertise of Ahmet Coskun. Coskun is a Georgia Tech immunoengineer known for his innovative approaches to understanding the immune response to cancer. An assistant professor in BME, Coskun holds the Bernie Marcus Early Career Professorship. He and his team use advanced imaging techniques and engineering principles to analyze tumor microenvironments in unprecedented detail. By examining how different immune cells interact with cancer cells, they hope to uncover the complexities of tumor biology and identify factors that contribute to treatment resistance.
This five-year, multi-million-dollar LLS SCOR award is the culmination of years of collaboration among leading researchers in the field of lymphoma. Singh, with colleagues Koff, Coskun, Christopher Flowers at MD Anderson Cancer Center, and Cornell Medicine’s Ari Melnick, Ethel Cesarman, and Leandro Cerchietti, are fostering a partnership in lymphomas and EBV-related cancers, which is instrumental in advancing research on lymphoma treatment health disparities. Their longstanding partnership reflects a commitment to addressing the complex challenges different populations face when battling deadly cancers.
"With this unique partnership, leveraging new cancer technologies, biology, and clinical expertise, we hope to make breakthroughs in lymphoma research and begin to address health disparities in lymphoma at multiscale levels,” said Melnick, a co-lead for LLS SCOR and Gebroe Family Professor of Hematology and Oncology at New York’s Weill Cornell Medicine.
The group also played a significant role in organizing, moderating, and presenting at the inaugural conference “Health Disparities in Hematologic Malignancies: From Genes to Outreach,” held in May 2023 in New York. The conference served as a vital platform for discussing the latest research, sharing best practices, and highlighting the importance of outreach initiatives aimed at improving care for underserved populations.
"The research will provide a unique window into the intricate structure of lymphomas and how these complexities influence treatment,” said Flowers, a physician-scientist and division head of Cancer Medicine at MD Anderson Cancer Center in Houston, Texas. “By studying lymphoma microenvironments in patient tissues and organoids, we can begin addressing health disparities in lymphoma, identifying why certain populations may respond differently to therapies. No other technology currently provides this level of insight or potential for tailored patient care."
This unique research collaboration is crucial, as understanding tumor heterogeneity can inform the development of more personalized treatment strategies, particularly for underserved communities that often face disparities in cancer care. By integrating engineering with oncology, the team hopes to create more effective therapies tailored to individual patient profiles, ultimately aiming to improve outcomes for all lymphoma patients. This multi-site collaboration aims to fast-track the development of therapies against lymphomas in African Americans and individuals with EBV-related conditions and eventually bring them to clinical trials.
Project Title: Translating molecular profiles into treatment approaches to target disparities in lymphoma
(Funding and award period: $5 million, October 1, 2024 - September 30, 2029)
News Contact
By: Savannah Williamson
Nov. 21, 2024
As Ahmet F. Coskun and his team of researchers continue their mission to create a 3D atlas of the human body, mapping cells and tissues, they’re making discoveries that could lead to better treatments for the most common type of lung cancer.
While they’re at it, they’re pioneering new fields of research, and possibly spinning the work into a new commercial venture.
Last year, Coskun and his team introduced a new study in “single cell spatial metabolomics,” which explores the distribution of small molecules — metabolites — within tissues and organs. Now they’re spearheading “spatial interactomics,” a research area concerned with interactions between various biomolecules inside of individual cells.
To study these interactions, they’ve developed an innovative technique, or tool, to better understand why non-small cell lung cancer, or NSCLC, resists treatment in so many patients. They call it the “intelligent sequential proximity ligation assay,” or iseqPLA.
“It’s a smart test that can look at proteins and how they interact with each other in space,” said Coskun, Bernie Marcus Early Career Professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University.
“Basically, we’re the first to create a new research area on spatial protein-protein interactions, which can tell us more about cell types and their functions,” said Coskun. “With spatial interactomics, we can validate how cells physically touch, sense, and regulate nearby cells through the interaction of pairs of proteins.”
So, the immediate goal of spatial interactomics is to investigate how protein-protein interactions drive drug resistance in NSCLC. And iseqPLA allows researchers to visualize how it’s all happening at the subcellular level. Coskun’s team described its work recently in the journal Nature Biomedical Engineering. He’s also forming a company to commercialize the technology.
Smarter Tools
Drugs called tyrosine kinase inhibitors (TKIs, like Osimertinib) have been successful in treating people with NSCLC. But many patients who initially respond well to the regimen, eventually develop a resistance. Protein interactions, a molecular kind of crosstalk, are a prime suspect in causing this resistance.
Proteins interact with each other all the time, and this mingling controls how cells grow, divide, or survive. Coskun and his team want to see how these interactions change in response to cancer treatment, and iseqPLA shows them, essentially attaching glowing tags to proteins, lighting up their locations and interactions under a microscope.
“Think of it like a super detailed map showing how different proteins in a cell are connected,” Coskun said.
The iseqPLA can examine 47 protein interactions in a single sample, which saves a lot of time (and resources) when compared to older methods, which look at two to three interactions at a time.
The researchers also created a computer model to analyze the spatial data they collected from iseqPLA, identifying patterns in protein interactions to help predict whether a cell was responding to a treatment or developing resistance.
“We showed that the test works not only in lab-grown cells but also in tissues from mice and humans,” Coskun said. “It can really help us understand how patients respond to certain treatments.”
Building a Spatial Omics Market
Going forward, Coskun aims to enhance iseqPLA to study interactions among RNA, proteins, and metabolites, as well as the RNA, proteins, metabolites, etc., and other subcellular dynamics. He also hopes to get the technology into the hands of other researchers.
“We believe it will be a groundbreaking tool,” he said.
With that in mind, Coskun is planning to form a startup company called SpatAllize. He’s working with VentureLab, the nonprofit organization at Georgia Tech that provides entrepreneurship programs for students and faculty.
“We are currently performing customer interviews and forming a strategy for a viable plan towards the marketplace,” he said.
He also plans to expand iseqPLA’s utility into other areas of research, focusing on how protein interactions influence the immune system, the heart, and brain health. His team is also developing a spatial interactomics robot that integrates iseqPLA with advanced imaging and automated deep learning.
“This will allow us to map all molecules within cells and tissues for an even better understanding of drug-cell interactions, particularly in cancer treatment planning,” Coskun said.
CITATION: Shuangyi Cai, Thomas Hu, Abhijeet Venkataraman, Felix G. Rivera Moctezuma, Efe Ozturk, Nicholas Zhang, Mingshuang Wang, Tatenda Zvidzai, Sandip Das, Adithya Pillai, Frank Schneider, Suresh S. Ramalingam, YouTake Oh, Shi-Yong Sun, and Ahmet F. Coskun. “Spatially resolved subcellular protein–protein interactomics in drug-perturbed lung-cancer cultures and tissues.” Nature Biomedical Engineering.
https://doi.org/10.1038/s41551-024-01271-x
FUNDING: This research was supported by the National Institutes of Health, grant Nos. P50CA217691, P30CA138292, and R33CA291197; and the National Science Foundation, grant No. R35GM151028. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of any funding agency.
COMPETING INTERESTS: Coskun, Cai, and Hu declare a patent application related to the spatial-signaling interactomics assay (U.S. Provisional 63/399,427 and U.S. Application No. 18/452,178).
News Contact
Pagination
- Previous page
- Page 13
- Next page