Apr. 01, 2026
In recent years, the Centers for Disease Control and Prevention, the Department of Homeland Security, and other authorities have flagged a record number of unauthorized shipments of biological materials. At the same time, global intelligence communities have identified numerous attempts to smuggle sensitive biological samples in efforts of industrial theft or espionage.
“A small vial of genetically engineered cells can contain multiple millions of dollars’ worth of intellectual property and require several years of work to develop,” said Corey Wilson, a professor in Georgia Tech’s School of Chemical and Biomolecular Engineering (ChBE). “Accordingly, the protection of high-value engineered cell lines has become critically important to the biotechnology industry.”
Wilson and his research team have published their findings in Science Advances demonstrating the effectiveness of their new biological security technology, known as GeneLock™, in protecting high-value engineered cell lines.
GeneLock is a cybersecurity-inspired technology that protects valuable genetic material directly at the DNA level. To demonstrate its strength, Wilson’s team conducted what they describe as a first-of-its-kind biohackathon, detailed in the new paper, to simulate unauthorized access.
“GeneLock greatly improves our ability to protect high-value engineered cell lines by expanding security from the lab environment to the genetic level,” Wilson said.
Economic Impact
What are the stakes? Estimates place the global market for high-value genetic materials at more than $1.5 trillion, projected to reach $8 trillion by 2035. The use of these materials ranges from advanced medicines and proprietary research enzymes to specialty chemicals and sustainable materials.
Currently, the protection of high-value cell lines depends on physical safeguards such as restricted lab access and secure facilities, Wilson explained.
“The key weakness of physical security measures is once circumvented, there are typically no measures in place to protect valuable cells from theft, abuse, or unauthorized use,” Wilson said.
“Once a sample leaves the building, the DNA it carries typically remains fully functional. This is like placing an unlocked cellphone in a desk drawer. Anyone who gains access to the drawer can view sensitive content on the phone—or in this case will have full access to the valuable cell line.”
Genetic Passcode Protection
The GeneLock biological security technology developed by Wilson and his team places a passcode on engineered cells, akin to those used on ATM machines and protected cellphones.
Instead of leaving a valuable gene in readable form, the team scrambles the DNA sequence of interest. The scrambled genetic asset remains in a nonfunctional state unless the living cell where it resides receives the correct sequence of chemical inputs. Those inputs act as a molecular passcode.
“Only the right combination, delivered in the right order, rearranges the DNA into a working form,” Wilson said.
Biohackathon Security Test
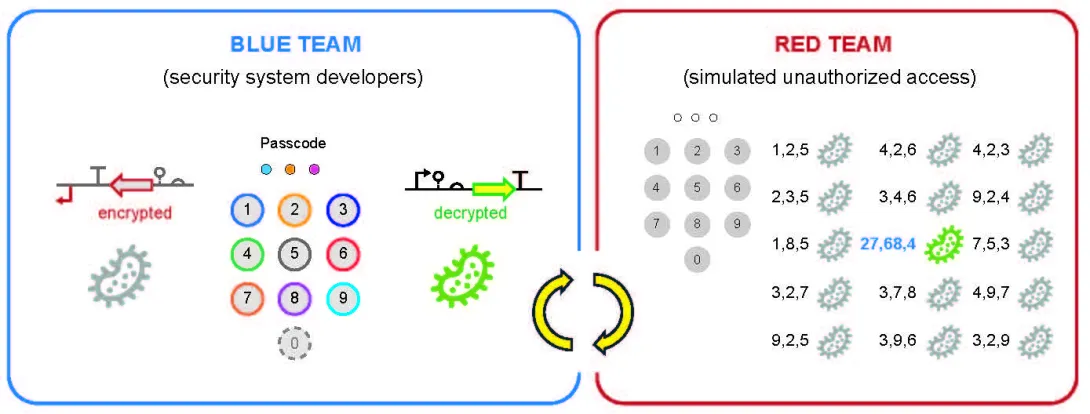
To evaluate the technology, the researchers organized a blue team and a red team in what they describe as an ethical biohackathon. The blue team designed the encrypted DNA sequence, while the red team was challenged to discover the correct chemical passcode through experimentation in a gray box exercise, meaning the red team had partial knowledge of the system but did not have access to the internal designs.
“This approach for testing security strength is commonly used in cybersecurity,” Wilson explained.
The blue team engineered the system inside Escherichia coli, or E. coli, a bacterium widely used in biotechnology. The protected asset was a fluorescent protein gene selected as a measurable stand-in for commercially valuable targets. When the correct chemical sequence was applied, the fluorescence turned on. Without the correct passcode, the gene remained scrambled and the cells could not fluoresce green.
“In practice, most DNA sequences produce valuable proteins or chemicals that are essentially invisible to the human eye, requiring specialized devices or experiments to observe,” Wilson said. “If the biohackathon were conducted with a standard commercially valuable target, the penetration testing would have taken more than 10 times longer to complete, years instead of months.”
The biohackathon results showed a dramatic reduction in risk. GeneLock reduced the probability of unlocking the genetic asset by random search to about 1 in 85,000 (a 0.001% chance), assuming the unauthorized user had access to the required chemical inputs.
Without access to those inputs, “the likelihood of success by chance becomes effectively negligible,” said Dowan Kim (Georgia Tech PhD 2024), co-lead author of the study.
Commercial Uses and What’s Next
Although the researchers used a non-commercial fluorescent protein as a test case, the implications extend much further. Many biotechnology companies rely on proprietary engineered strains. New England Biolabs, for example, produces more than 265 non-disclosed enzymes in E. coli, each representing a high-value cell line.
Protein-based drugs are also manufactured in living cells, and proprietary metabolic pathways are used to produce specialty chemicals, bioplastics, and high-value ingredients.
“In each case, the genetic blueprint inside the cell represents intellectual property that can be protected by our technology,” said Ishita Kumar, a PhD candidate in ChBE and co-lead author of the study.
While the team’s current focus is on protecting intellectual property in the form of high-value cells, future iterations aim to strengthen biological security more broadly.
“We are currently developing protection measures to mitigate unauthorized use or release of sensitive cell lines that can be potentially hazardous to human health or the environment,” Wilson said.
“As it stands, GeneLock represents an important shift in biological security, enabling, for the first time, protection of valuable cells at the genetic level, even after physical security measures have been bypassed,” he added.
The work is already moving toward commercialization. The team filed a provisional patent application with the U.S. Patent and Trademark Office in February 2026 and is forming a company to deploy the technology.
This research was funded by a grant from the National Science Foundation.
CITATION:
Dowan Kim, Ishita Kumar, Mohamed Hassan, Luisa F. Barraza-Vergara, Christopher A. Voigt, and Corey J. Wilson, “Protecting cells at the genetic level and simulating unauthorized access via a biohackathon,” Science Advances, 2026.
News Contact
Brad Dixon, braddixon@gatech.edu
Dec. 04, 2011
When RNA component units called ribonucleotides become embedded in genomic DNA, which contains the complete genetic data for an organism, they can cause problems for cells. It is known that ribonucleotides in DNA can potentially distort the DNA double helix, resulting in genomic instability and altered DNA metabolism, but not much is known about the fate of these ribonucleotides.
A new study provides a mechanistic explanation of how ribonucleotides embedded in genomic DNA are recognized and removed from cells. Two mechanisms, enzymes called ribonucleases (RNases) H and the DNA mismatch repair system, appear to interplay to root out the RNA components.
"We believe this is the first study to show that cells utilize independent repair pathways to remove mispaired ribonucleotides embedded in chromosomal DNA, which can be sources of genetic modification if not removed," said Francesca Storici, an assistant professor in the School of Biology at the Georgia Institute of Technology. "The results also highlight a novel case of genetic redundancy, where the mismatch repair system and RNase H mechanisms compete with each other to remove misincorporated ribonucleotides and restore DNA integrity."
The findings were reported Dec. 4, 2011 in the advance online publication of the journal Nature Structural & Molecular Biology. The research was supported by the Georgia Cancer Coalition, National Science Foundation and Georgia Tech Integrative BioSystems Institute.
Storici and Georgia Tech biology graduate students Ying Shen and Kyung Duk Koh conducted the study in collaboration with Bernard Weiss, a professor emeritus in the Department of Pathology and Laboratory Medicine at Emory University.
"We wanted to understand how cells of the bacterium Escherichia coli and the yeast Saccharomyces cerevisiae tolerate the presence of different ribonucleotides embedded in their genomic DNA. We found that the structure of a ribonucleotide tract embedded in DNA influenced its ability to cause genetic mutations more than the tract's length," said Storici.
With double-stranded DNA, when wrong bases are paired or one or few nucleotides are in excess or missing on one of the strands, a mismatch is generated. If mismatches are not corrected, they can lead to mutations.
The researchers found that single mismatched ribonucleotides in chromosomal DNA were removed by either the mismatch repair system or RNase H type 2. Mismatched ribonucleotides in the middle of at least four other ribonucleotides required RNase H type 1 for removal.
"We were excited to find that a DNA repair mechanism like mismatch repair was activated by RNA/DNA mismatches and could remove ribonucleotides embedded in chromosomal DNA," explained Storici. "In future studies, we plan to test whether other DNA repair mechanisms, such as nucleotide-excision repair and base-excision repair, can also locate and remove ribonucleotides in DNA."
Using gene correction assays driven by short nucleic acid polymers called oligonucleotides, the researchers showed that when ribonucleotides embedded in DNA were not removed, they served as templates for DNA synthesis and produced a mutation in the DNA. If both the mismatch repair system and RNase H repair mechanisms are disabled, ribonucleotide-driven gene modification increased by a factor of 47 in the yeast and 77,000 in the bacterium.
Defects in the mismatch repair system are known to predispose a person to certain types of cancer. Because the mismatch repair system is conserved from unicellular to multicellular organisms, such as humans, this study's findings open up the possibility that defects in the mismatch repair system could have consequences more critical than previously thought given the newly identified function of mismatch repair to target RNA/DNA mispairs.
The results also provide new information on the capacity of RNA to play an active role in DNA editing and remodeling, which could be the basis of an unexplored process of RNA-driven DNA evolution.
This project was supported by the National Science Foundation (NSF) (Award No. MCB-1021763). The content is solely the responsibility of the principal investigators and does not necessarily represent the official views of the NSF.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364