May. 06, 2012
Gaining access to the inner workings of a neuron in the living brain offers a wealth of useful information: its patterns of electrical activity, its shape, even a profile of which genes are turned on at a given moment. However, achieving this entry is such a painstaking task that it is considered an art form; it is so difficult to learn that only a small number of labs in the world practice it.
But that could soon change: Researchers at MIT and the Georgia Institute of Technology have developed a way to automate the process of finding and recording information from neurons in the living brain. The researchers have shown that a robotic arm guided by a cell-detecting computer algorithm can identify and record from neurons in the living mouse brain with better accuracy and speed than a human experimenter.
The new automated process eliminates the need for months of training and provides long-sought information about living cells’ activities. Using this technique, scientists could classify the thousands of different types of cells in the brain, map how they connect to each other, and figure out how diseased cells differ from normal cells.
The project is a collaboration between the labs of Ed Boyden, associate professor of biological engineering and brain and cognitive sciences at MIT, and Craig Forest, an assistant professor in the George W. Woodruff School of Mechanical Engineering at Georgia Tech.
“Our team has been interdisciplinary from the beginning, and this has enabled us to bring the principles of precision machine design to bear upon the study of the living brain,” Forest says. His graduate student, Suhasa Kodandaramaiah, spent the past two years as a visiting student at MIT, and is the lead author of the study, which appears in the May 6 issue of Nature Methods.
The method could be particularly useful in studying brain disorders such as schizophrenia, Parkinson’s disease, autism and epilepsy, Boyden says. “In all these cases, a molecular description of a cell that is integrated with [its] electrical and circuit properties … has remained elusive,” says Boyden, who is a member of MIT’s Media Lab and McGovern Institute for Brain Research. “If we could really describe how diseases change molecules in specific cells within the living brain, it might enable better drug targets to be found.”
Automation
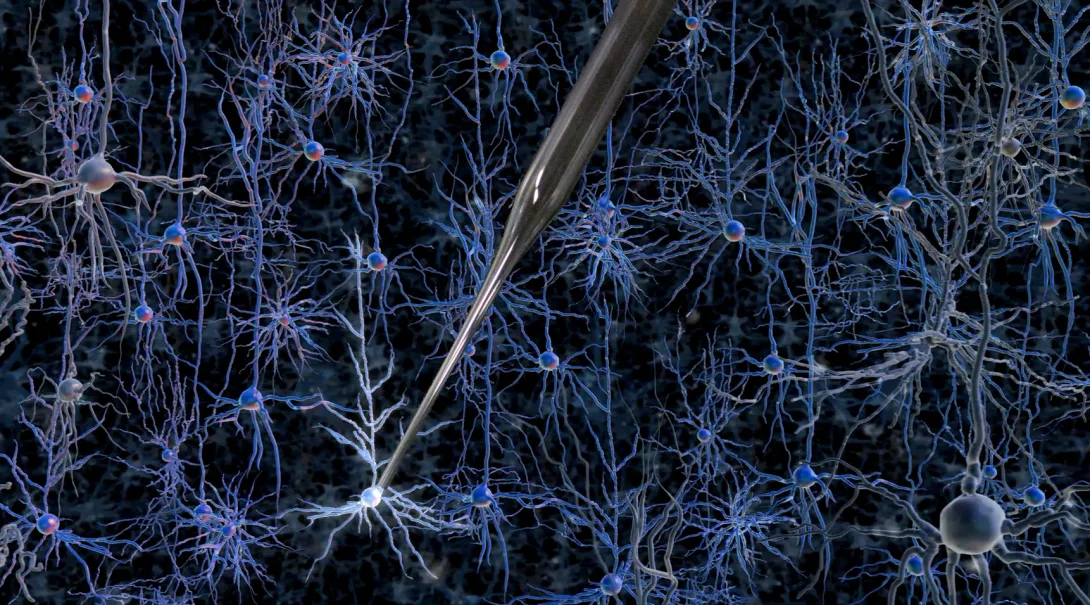
Kodandaramaiah, Boyden and Forest set out to automate a 30-year-old technique known as whole-cell patch clamping, which involves bringing a tiny hollow glass pipette in contact with the cell membrane of a neuron, then opening up a small pore in the membrane to record the electrical activity within the cell. This skill usually takes a graduate student or postdoc several months to learn.
Kodandaramaiah spent about four months learning the manual patch-clamp technique, giving him an appreciation for its difficulty. “When I got reasonably good at it, I could sense that even though it is an art form, it can be reduced to a set of stereotyped tasks and decisions that could be executed by a robot,” he says.
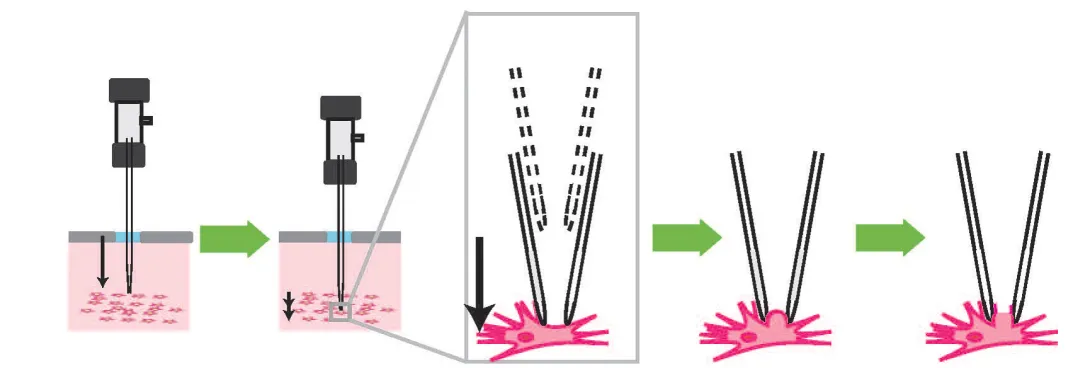
To that end, Kodandaramaiah and his colleagues built a robotic arm that lowers a glass pipette into the brain of an anesthetized mouse with micrometer accuracy. As it moves, the pipette monitors a property called electrical impedance — a measure of how difficult it is for electricity to flow out of the pipette. If there are no cells around, electricity flows and impedance is low. When the tip hits a cell, electricity can’t flow as well and impedance goes up.
The pipette takes two-micrometer steps, measuring impedance 10 times per second. Once it detects a cell, it can stop instantly, preventing it from poking through the membrane. “This is something a robot can do that a human can’t,” Boyden says.
Once the pipette finds a cell, it applies suction to form a seal with the cell’s membrane. Then, the electrode can break through the membrane to record the cell’s internal electrical activity. The robotic system can detect cells with 90 percent accuracy, and establish a connection with the detected cells about 40 percent of the time.
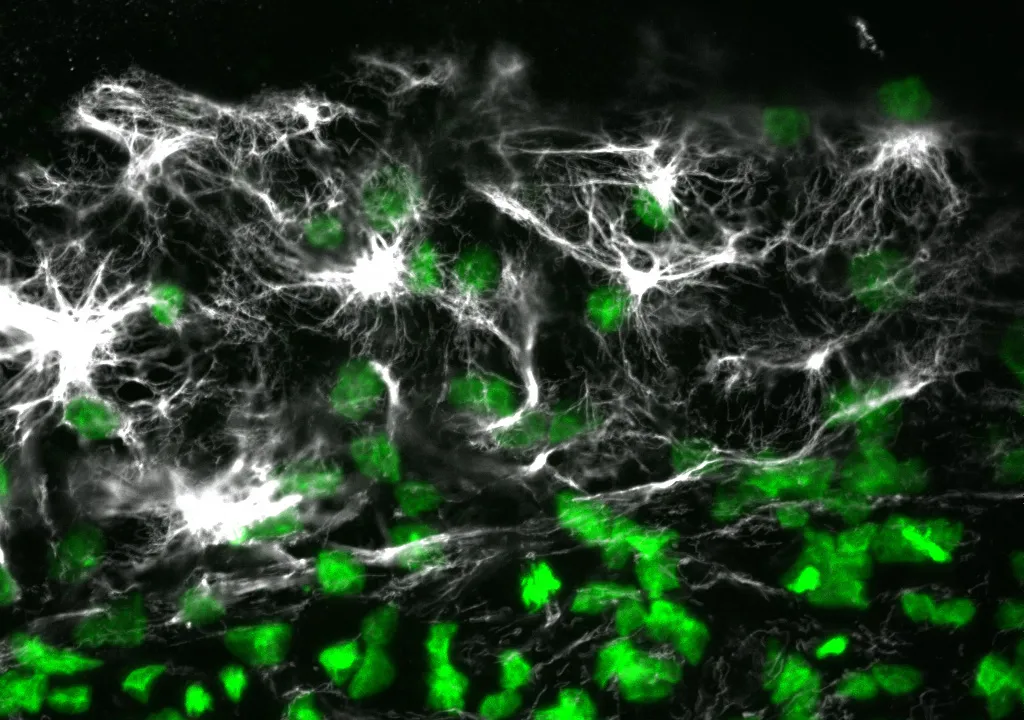
The researchers also showed that their method can be used to determine the shape of the cell by injecting a dye; they are now working on extracting a cell’s contents to read its genetic profile.
Development of the new technology was funded primarily by the National Institutes of Health, the National Science Foundation and the MIT Media Lab.
New era for robotics
The researchers recently created a startup company, Neuromatic Devices, to commercialize the device.
The researchers are now working on scaling up the number of electrodes so they can record from multiple neurons at a time, potentially allowing them to determine how different parts of the brain are connected.
They are also working with collaborators to start classifying the thousands of types of neurons found in the brain. This “parts list” for the brain would identify neurons not only by their shape — which is the most common means of classification — but also by their electrical activity and genetic profile.
“If you really want to know what a neuron is, you can look at the shape, and you can look at how it fires. Then, if you pull out the genetic information, you can really know what’s going on,” Forest says. “Now you know everything. That’s the whole picture.”
Boyden says he believes this is just the beginning of using robotics in neuroscience to study living animals. A robot like this could potentially be used to infuse drugs at targeted points in the brain, or to deliver gene therapy vectors. He hopes it will also inspire neuroscientists to pursue other kinds of robotic automation — such as in optogenetics, the use of light to perturb targeted neural circuits and determine the causal role that neurons play in brain functions.
Neuroscience is one of the few areas of biology in which robots have yet to make a big impact, Boyden says. “The genome project was done by humans and a giant set of robots that would do all the genome sequencing. In directed evolution or in synthetic biology, robots do a lot of the molecular biology,” he says. “In other parts of biology, robots are essential.”
Other co-authors include MIT grad student Giovanni Talei Franzesi and MIT postdoc Brian Y. Chow.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or Caroline McCall (cmccall5@mit.edu; 617-253-1682)
Writer: Anne Trafton, MIT News
News Contact
Abby Robinson
Research News and Publications
abby@innovate.gatech.edu
404-385-3364
Feb. 09, 2012
Recent earthquake damage has exposed the vulnerability ofexisting structures to strong ground movement. At the Georgia Institute ofTechnology, researchers are analyzing shape-memory alloys for their potentialuse in constructing seismic-resistant structures.
“Shape-memory alloys exhibit unique characteristics that youwould want for earthquake-resistant building and bridge design and retrofitapplications: they have the ability to dissipate significant energy withoutsignificant degradation or permanent deformation,” said Reginald DesRoches, a professorin the School of Civil and Environmental Engineering at Georgia Tech.
Georgia Tech researchers have developed a model thatcombines thermodynamics and mechanical equations to assess what happens whenshape-memory alloys are subjected to loading from strong motion. The researchersare using the model to analyze how shape-memory alloys in a variety ofcomponents -- cables, bars, plates and helical springs -- respond to different loadingconditions. From that information, they can determine the optimalcharacteristics of the material for earthquake applications.
The model was developed by DesRoches, School of MechanicalEngineering graduate student Reza Mirzaeifar, School of Civil and EnvironmentalEngineering associate professor Arash Yavari, and School of Mechanical Engineeringand School of Materials Science and Engineering professor Ken Gall.
A paper describing the thermo-mechanical model was publishedonline Feb. 3 in the InternationalJournal of Non-Linear Mechanics. This research was supported by theTransportation Research Board IDEA program.
To improve the performance of structures during earthquakes,researchers around the world have been investigating the use of “smart”materials, such as shape-memory alloys, which can bounce back afterexperiencing large loads. The most common shape-memory alloys are made of metalmixtures containing copper-zinc-aluminum-nickel, copper-aluminum-nickel ornickel-titanium. Potential applications of shape-memory alloys in bridge andbuilding structures include their use in bearings, columns and beams, orconnecting elements between beams and columns. But before this class ofmaterials can be used, the effect of extreme and repetitive loads on thesematerials must be thoroughly examined.
“For standard civil engineering materials, you can usemechanics to look at force and displacement to measure stress and strain, butfor this class of shape-memory alloys that changes properties when it undergoesloading and unloading, you have to consider thermodynamics and mechanics,” explainedYavari.
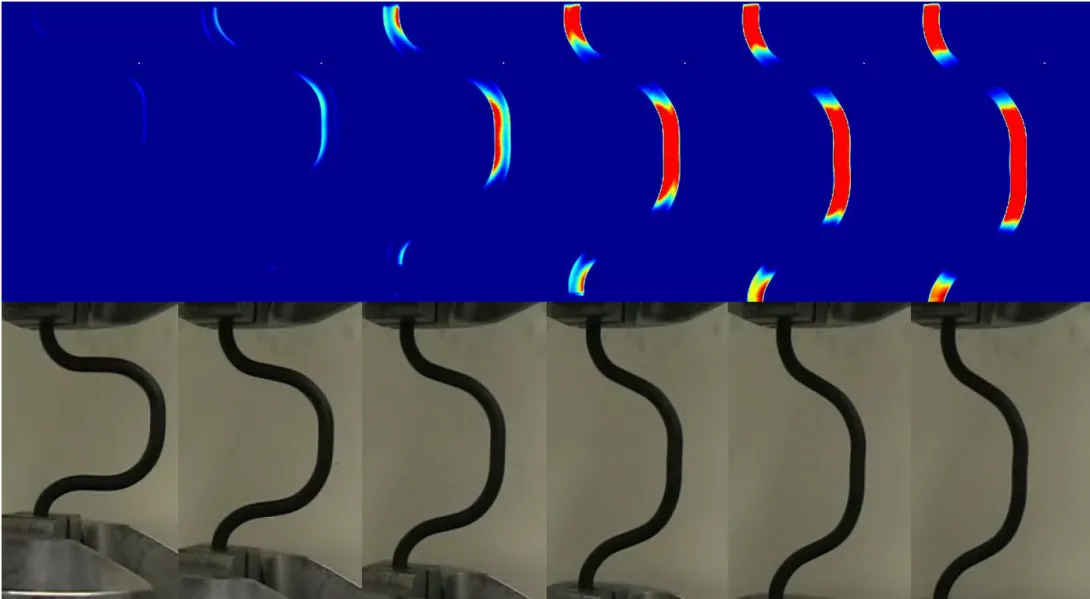
The Georgia Tech team found that the generation andabsorption of heat during loading and unloading caused a temperature gradientin shape-memory alloys, which caused a non-uniform stress distribution in thematerial even when the strain was uniform.
“Shape-memory alloys previously examined in detail werereally thin wires, which can exchange heat with the ambient environment rapidlyand no temperature change is seen,” said Mirzaeifar. “When you start to examinealloys in components large enough to be used in civil engineering applications,the internal temperature is no longer uniform and needs to be taken intoaccount.”
To predict the internal temperature distribution ofshape-memory alloys under loading-unloading cycles, which could then be used todetermine the stress distribution, the researchers developed a model that usedthe surface thermal boundary conditions, diameter and loading rate of the alloyas inputs.
The team included ambient conditions in the model becauseshape-memory alloys for seismic applications could operate in a variety ofenvironments -- such as water if used in bridge structures or air if used inbuilding structures -- which would produce different rates of heat transfer. Theresearchers used a thermal camera to record the variation in surfacetemperature of shape-memory alloys experiencing loading and unloading.
Using their model, the researchers were able to accuratelypredict internal temperature and stress distributions for shape-memory alloys. Themodel results were verified with experimental tests. In one test, they foundthat a shape-memory alloy loaded at a very slow rate had time to exchange theheat created with the ambient environment and exhibited uniform stress. If it wasloaded very rapidly, it did not have enough time to exchange the heat, leadingto a non-uniform stress distribution.
“Our analytical solutions are exact, fast and capable of simulatingthe complicated coupled thermo-mechanical response of shape-memory alloysconsidering temperature changes and loading rate dependency,” said Mirzaeifar.
In future work, the researchers plan to examine morecomplicated shapes and the effects of combination loading -- tension, bendingand torsion -- to optimize shape-memory alloys for earthquake applications.
This project issupported by the Transportation Research Board of the National Academies (AwardNo. NCHRP-147). The National Academies has rights to the data and the contentis solely the responsibility of the principal investigators and does notnecessarily represent the official views of the National Academies.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media RelationsContacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or JohnToon (jtoon@gatech.edu; 404-894-6986)
Writer: AbbyRobinson
News Contact
Abby Robinson
Research News and Publications
abby@innovate.gatech.edu
404-385-3364
Sep. 29, 2011
The National Science Foundation (NSF) has awarded $6 million to fund three projects involving researchers from the Georgia Institute of Technology. Each four-year, $2 million grant was awarded through the NSF's Division of Emerging Frontiers in Research and Innovation (EFRI).
"The EFRI research teams will probe some profound aspects of the interface of biology and engineering," said Sohi Rastegar, director of EFRI. "If they are successful, the principles and theories uncovered in their investigations could unlock many technological opportunities."
This year, 14 transformative, fundamental research projects were awarded EFRI grants in two emerging areas: technologies that build on understanding of biological signaling, and machines that can interact and cooperate with humans.
The three Georgia Tech projects include:
- Developing a "therapeutic robot" to help rehabilitate and improve motor skills in people with mobility problems;
- Creating wearable sensors that allow blind people to "see" with their hands, bodies or faces;
- Generating and rigorously testing quantitative models that describe spatial and temporal regulation of cell differentiation in tissues.
The therapeutic robot could enhance, assist and improve motor skills in humans with varying motor capabilities and deficits. The goal of the project is to program a humanoid rehabilitation robot to perform a "partnered box step," which is a defined pattern of weight shifts and directional changes, solely based on interpreting movement cues from subtle changes in forces between the hands and arms of the robot and the person.
To do this, researchers at Georgia Tech and Emory University will study how humans use their muscles to walk, balance and generate force signals with the hands for guidance when moving in cooperation with another person. They will also study "rehabilitative partnered dance," which has been specifically adapted to help improve gait and balance in individuals with motor impairments.
"Our vision is to develop robots that will interact with humans as both assistants and movement therapists," explained principal investigator Lena Ting, an associate professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. "We expect our project to have a long-term impact on quality of life of individuals with movement difficulties, such as those caused by Parkinson's disease, stroke and injury by improving fitness, motor skills and social engagement."
Working with Ting on the project are Emory University School of Medicine (geriatrics) assistant professor Madeleine Hackney, Coulter Department of Biomedical Engineering assistant professor Charlie Kemp and Georgia Tech School of Interactive Computing assistant professor Karen Liu.
For the second project, researchers at Georgia Tech and The City College of New York will investigate devices for "alternative perception" and the principles underlying the human-machine interaction. Alternative perception combines electronics and the other senses to emulate vision. In addition to aiding the visually impaired, the findings are expected to have other applications, such as the development of intelligent robots.
The researchers plan to untangle how humans learn to coordinate input from their senses -- e.g. vision, touch -- with movements, like reaching for a glass or moving through a crowded room. They will then map out how machines, such as robots and computers, learn similar tasks, to model devices that can assist humans.
The team envisions a multifunctional array of sensors on the body and has already developed prototypes for some of the devices. The full complement of wearable sensors would help a sightless person navigate by conveying information about his or her surroundings.
The researchers hope their findings on perception, and the prototypes they develop, will spawn a raft of wearable electronic devices to help blind people "see" their environment at a distance through touch, hearing and other senses. The technology would also benefit sighted individuals who must navigate in poor visibility, such as firefighters and pilots.
Principal investigator Zhigang Zhu, professor of computer science and computer engineering in City College's Grove School of Engineering, will collaborate with City College professor of psychology and director of the Program in Cognitive Neuroscience Tony Ro, City College professor of electrical engineering Ying Li Tian, Georgia Tech Woodruff School of Mechanical Engineering professor Kok-Meng Lee, and Georgia Tech School of Applied Physiology associate professor Boris Prilutsky.
The third project will address a fundamental question of developmental biology: what controls the spatial and temporal patterns of cell differentiation? Answering this question will lead to a better understanding of the basic principles of embryogenesis, explain origins of developmental disorders, and provide guidelines for tissue engineering and regenerative medicine.
The research will be conducted by principal investigator and Princeton University Department of Chemical and Biological Engineering associate professor Stanislav Shvartsman, Georgia Tech School of Chemical and Biomolecular Engineering associate professor Hang Lu, New York University Department of Biology professor Christine Rushlow, and University of Illinois at Urbana Champaign Department of Computer Science associate professor Saurabh Sinha.
Scientists know that among an embryo's first major developments is the establishment of its dorsoventral axis, which runs from its back to its belly. The researchers plan to study how this axis development unfolds -- specifically the presence and location of proteins during the process, which give rise to muscle, nerve and skin tissues.
To enable large-scale quantitative analyses of protein positional information along the dorsoventral axis, Lu and Shvartsman will further develop a microfluidic device they previously designed to reliably and robustly orient several hundred embryos in just a few minutes.
"By understanding this system at a deeper, quantitative level, we will elucidate general principles underlying the operation of genetic and multicellular networks that drive development," said Lu.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Mar. 23, 2011
Avian embryos could join the list of model organisms used to study a specific type of cell migration called epiboly, thanks to the results of a study published this month in the journal Developmental Dynamics. The new study provides insights into the mechanisms of epiboly, a developmental process involving mass movement of cells as a sheet, which is linked with medical conditions that include wound healing and cancer.
The study, published online on March 15, explains how epithelial cells expand as a sheet and migrate to engulf the entire avian egg yolk as it grows. It also reveals the presence of certain molecules during this process that have not been previously reported in other major developmental models, including Xenopus frogs and zebrafish.
"These molecules and mechanisms of early development in the avian embryo may demonstrate evolutionary differences across species in the collective movement of epithelial cells and motivate additional studies of avian embryo development," said Evan Zamir, an assistant professor in the George W. Woodruff School of Mechanical Engineering at Georgia Tech.
Matt Futterman, who worked on the project as a graduate student at Georgia Tech, and mechanical engineering professor Andrés García also contributed to this study. The research was funded by Zamir's new faculty support from Georgia Tech and by a grant to García from the National Institutes of Health.
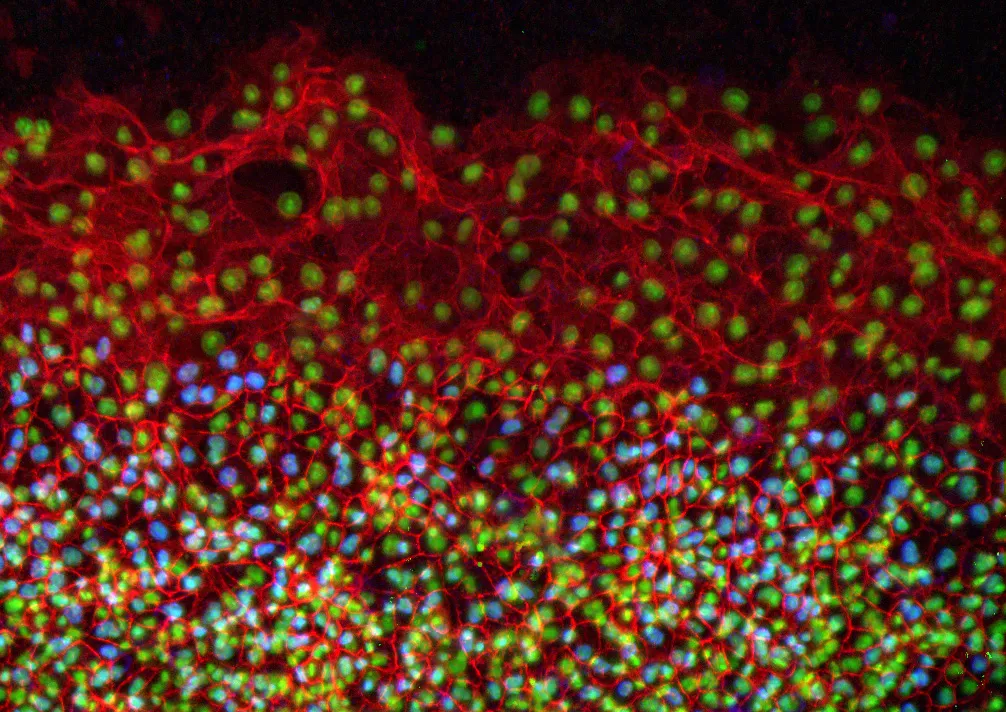
In the study, the researchers conducted immunofluorescence and high-resolution confocal microscopy experiments to examine the spatial distribution and expression of five proteins -- vimentin, cytokeratin, β-catenin, E-cadherin and laminin -- as cells moved to wrap the yolk sac of quail embryos during development.
The results showed that during this process, four of the proteins -- vimentin, cytokeratin, β-catenin and E-cadherin -- appeared in the cells located at the free edge of the migrating cell sheet. Finding dense interconnected networks of both vimentin and cytokeratin in the edge cells surprised the researchers.
"Since cytokeratin is generally associated with the epithelial phenotype and vimentin is generally associated with the mesenchymal phenotype, it's rare to see them expressed in the same cells, but this does occur in metastasizing tumor cells," said Zamir.
Cells expressing the mesenchymal phenotype are typically found in connective tissues -- such as bone, cartilage, and the lymphatic and circulatory systems -- whereas cells of the epithelial phenotype are found in cavities and glands and on surfaces throughout the body.
This finding provides evidence that epithelial cells normally attached to a membrane surface underwent biochemical changes that enabled them to assume a mesenchymal cell phenotype, which enhanced their migratory capacity. This process, called partial epithelial-to-mesenchymal transition, has many similarities to the initiation of tumor cell metastasis and wound healing.
Since this epithelial and mesenchymal expression pattern in the edge cells has not previously been reported in Xenopus or zebrafish, it may be unique to the avian embryo. This discovery would make the avian embryo a valuable model for studying tumor cell migration and wound healing.
In addition to detailing protein expression in the quail embryo during development, the researchers also determined the origin of the new cells required at the migrating edge to cover the growing yolk. During development, the radius of the quail yolk doubles every day for the first few days, representing a hundreds-fold increase in the egg yolk surface area.
"For each individual cell that has to cover the egg yolk as it grows, the migration around the yolk is extraordinary, because it's such a large territory -- it would be like an ant walking across the earth," explained Zamir.
Looking more closely at the edge cells, the researchers found strong evidence that expansion of the edge cell population was due exclusively to cells relocating from an interior region to the edge as the embryo expanded. The cells located at the free edge generated the bulk of the traction force necessary for expansion and towed the cells within the interior of the epithelium.
"These experiments confirm that edge cell proliferation is not the primary mechanism for expansion of the edge cell population," noted Zamir. "And our observation of epithelial-to-mesenchymal transition in the edge cells explains how these epithelial cells might be changing phenotype to become migratory in this rapidly expanding sheet."
To determine if this study's findings are indeed unique to the avian embryo, Zamir plans to conduct further studies to characterize protein expression and cell migration in Xenopus and zebrafish.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364