May. 10, 2012
New research findings show that embryonic stem cells unable to fully compact the DNA inside them cannot complete their primary task: differentiation into specific cell types that give rise to the various types of tissues and structures in the body.
Researchers from the Georgia Institute of Technology and Emory University found that chromatin compaction is required for proper embryonic stem cell differentiation to occur. Chromatin, which is composed of histone proteins and DNA, packages DNA into a smaller volume so that it fits inside a cell.
A study published on May 10, 2012 in the journal PLoS Genetics found that embryonic stem cells lacking several histone H1 subtypes and exhibiting reduced chromatin compaction suffered from impaired differentiation under multiple scenarios and demonstrated inefficiency in silencing genes that must be suppressed to induce differentiation.
“While researchers have observed that embryonic stem cells exhibit a relaxed, open chromatin structure and differentiated cells exhibit a compact chromatin structure, our study is the first to show that this compaction is not a mere consequence of the differentiation process but is instead a necessity for differentiation to proceed normally,” said Yuhong Fan, an assistant professor in the Georgia Tech School of Biology.
Fan and Todd McDevitt, an associate professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University, led the study with assistance from Georgia Tech graduate students Yunzhe Zhang and Kaixiang Cao, research technician Marissa Cooke, and postdoctoral fellow Shiraj Panjwani.
The work was supported by the National Institutes of Health’s National Institute of General Medical Sciences (NIGMS), the National Science Foundation, a Georgia Cancer Coalition Distinguished Scholar Award, and a Johnson & Johnson/Georgia Tech Healthcare Innovation Award.
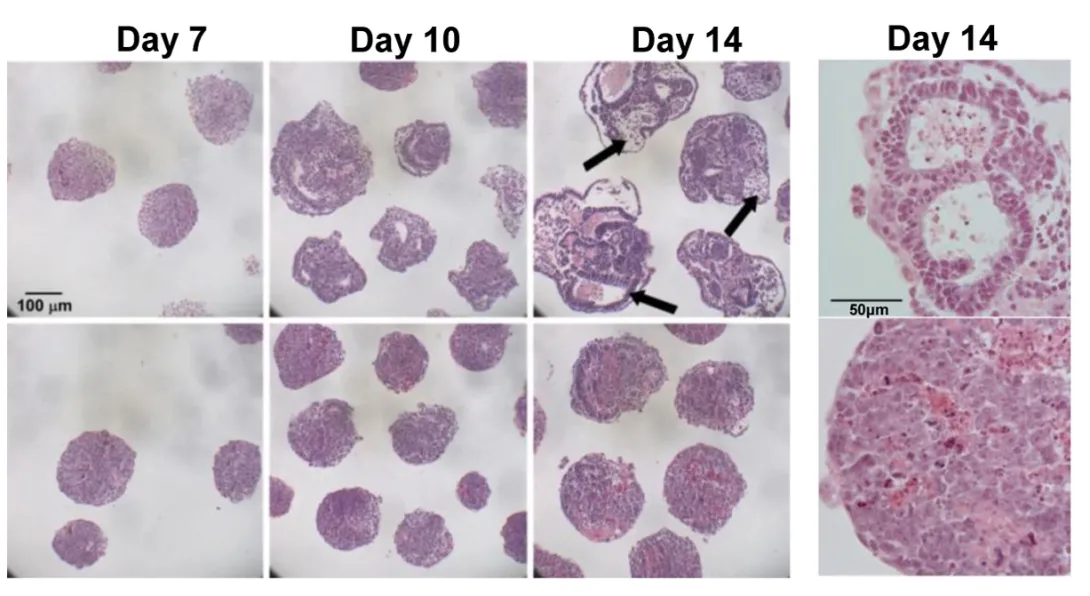
To investigate the impact of linker histones and chromatin folding on stem cell differentiation, the researchers used embryonic stem cells that lacked three subtypes of linker histone H1 -- H1c, H1d and H1e -- which is the structural protein that facilitates the folding of chromatin into a higher-order structure. They found that the expression levels of these H1 subtypes increased during embryonic stem cell differentiation, and embryonic stem cells lacking these H1s resisted spontaneous differentiation for a prolonged time, showed impairment during embryoid body differentiation and were unsuccessful in forming a high-quality network of neural cells.
“This study has uncovered a new, regulatory function for histone H1, a protein known mostly for its role as a structural component of chromosomes,” said Anthony Carter, who oversees epigenetics grants at NIGMS. “By showing that H1 plays a part in controlling genes that direct embryonic stem cell differentiation, the study expands our understanding of H1’s function and offers valuable new insights into the cellular processes that induce stem cells to change into specific cell types.”
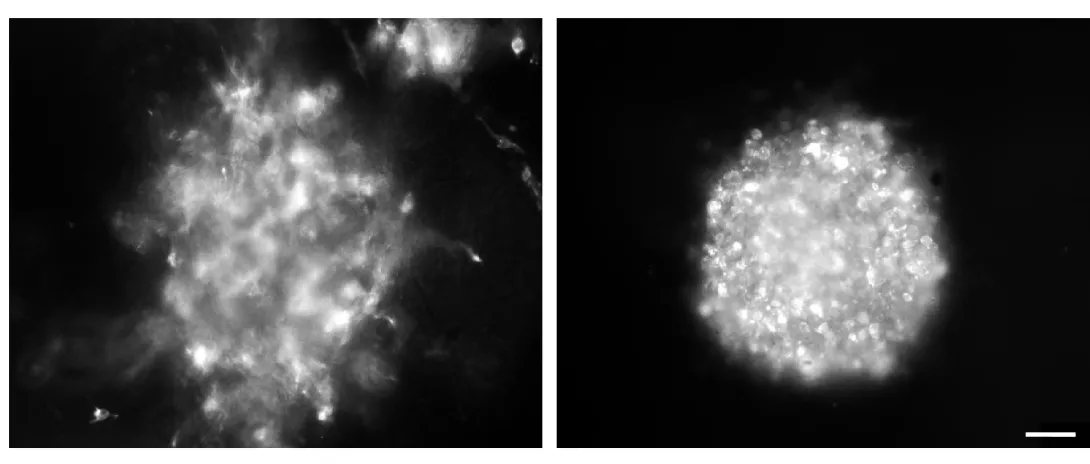
During spontaneous differentiation, the majority of the H1 triple-knockout embryonic stem cells studied by the researchers retained a tightly packed colony structure typical of undifferentiated cells and expressed high levels of Oct4 for a prolonged time. Oct4 is a pluripotency gene that maintains an embryonic stem cell’s ability to self-renew and must be suppressed to induce differentiation.
“H1 depletion impaired the suppression of the Oct4 and Nanog pluripotency genes, suggesting a novel mechanistic link by which H1 and chromatin compaction may mediate pluripotent stem cell differentiation by contributing to the epigenetic silencing of pluripotency genes,” explained Fan. “While a significant reduction in H1 levels does not interfere with embryonic stem cell self-renewal, it appears to impair differentiation.”
The researchers also used a rotary suspension culture method developed by McDevitt to produce with high efficiency homogonous 3D clumps of embryonic stem cells called embryoid bodies. Embryoid bodies typically contain cell types from all three germ layers -- the ectoderm, mesoderm and endoderm -- that give rise to the various types of tissues and structures in the body. However, the majority of the H1 triple-knockout embryoid bodies formed in rotary suspension culture lacked differentiated structures and displayed gene expression signatures characteristic of undifferentiated stem cells.
“H1 triple-knockout embryoid bodies displayed a reduced level of activation of many developmental genes and markers in rotary culture, suggesting that differentiation to all three germ layers was affected.” noted McDevitt.
The embryoid bodies also lacked the epigentic changes at the pluripotency genes necessary for differentiation, according to Fan.
“When we added one of the deleted H1 subtypes to the embryoid bodies, Oct4 was suppressed normally and embryoid body differentiation continued,” explained Fan. “The epigenetic regulation of Oct4 expression by H1 was also evident in mouse embryos.”
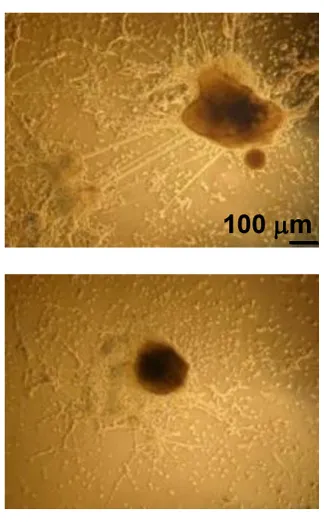
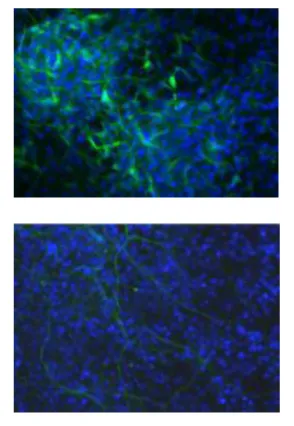
In another experiment, the researchers provided an environment that would encourage embryonic stem cells to differentiate into neural cells. However, the H1 triple-knockout cells were defective in forming neuronal and glial cells and a neural network, which is essential for nervous system development. Only 10 percent of the H1 triple-knockout embryoid bodies formed neurites and they produced on average eight neurites each. In contrast, half of the normal embryoid bodies produced, on average, 18 neurites.
In future work, the researchers plan to investigate whether controlling H1 histone levels can be used to influence the reprogramming of adult cells to obtain induced pluripotent stem cells, which are capable of differentiating into tissues in a way similar to embryonic stem cells.
Research reported in this publication was supported by the National Institute of General Medical Sciences of the National Institutes of Health (NIH) under award number GM085261 and the National Science Foundation under award number CBET-0939511. The content is solely the responsibility of the principal investigators and does not necessarily represent the official views of the NIH or NSF.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
abby@innovate.gatech.edu
404-385-3364
Mar. 28, 2012
Treating invasive brain tumors with a combination of chemotherapy and radiation has improved clinical outcomes, but few patients survive longer than two years after diagnosis. The effectiveness of the treatment is limited by the tumor’s aggressive invasion of healthy brain tissue, which restricts chemotherapy access to the cancer cells and complicates surgical removal of the tumor.
To address this challenge, researchers from the Georgia Institute of Technology and Emory University have designed a new treatment approach that appears to halt the spread of cancer cells into normal brain tissue in animal models. The researchers treated animals possessing an invasive tumor with a vesicle carrying a molecule called imipramine blue, followed by conventional doxorubicin chemotherapy. The tumors ceased their invasion of healthy tissue and the animals survived longer than animals treated with chemotherapy alone.
“Our results show that imipramine blue stops tumor invasion into healthy tissue and enhances the efficacy of chemotherapy, which suggests that chemotherapy may be more effective when the target is stationary,” said Ravi Bellamkonda, a professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. “These results reveal a new strategy for treating brain cancer that could improve clinical outcomes.”
The results of this work were published on March 28, 2012 in the journal Science Translational Medicine. The research was supported primarily by the Ian’s Friends Foundation and partially by the Georgia Cancer Coalition, the Wallace H. Coulter Foundation and a National Science Foundation graduate research fellowship.
In addition to Bellamkonda, collaborators on the project include Jack Arbiser, a professor in the Emory University Department of Dermatology; Daniel Brat, a professor in the Emory University Department of Pathology and Laboratory Medicine; and the paper’s lead author, Jennifer Munson, a former Fulbright Scholar who was a bioengineering graduate student in the Georgia Tech School of Chemical & Biomolecular Engineering when the research was conducted.
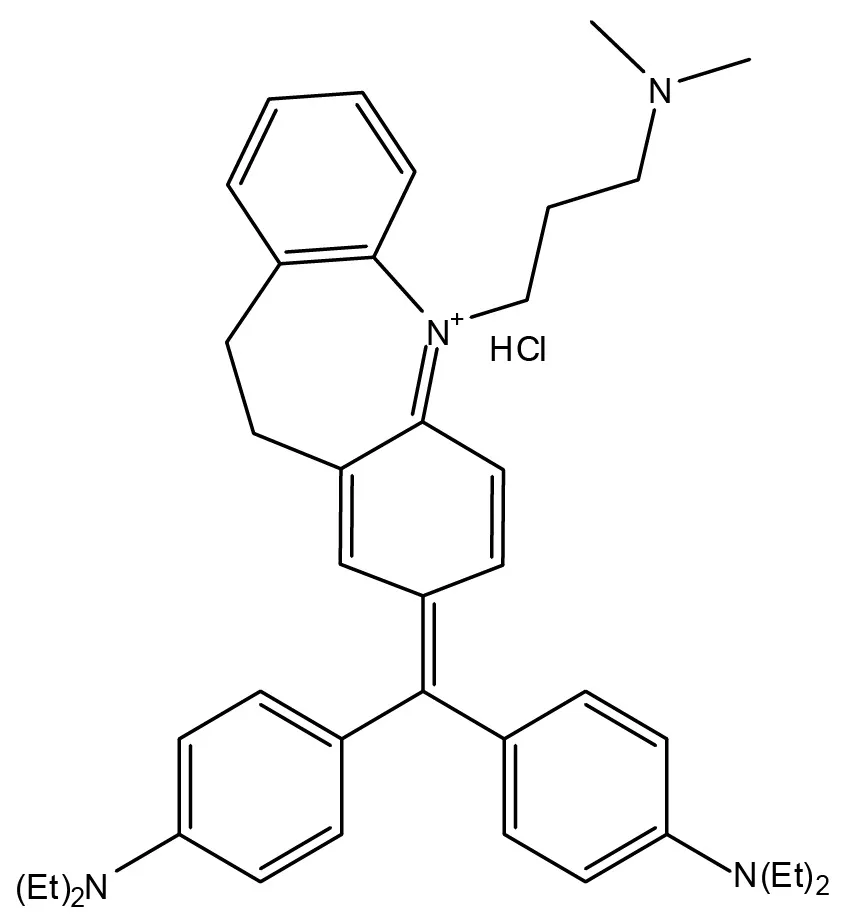
Arbiser designed the novel imipramine blue compound, which is an organic triphenylmethane dye. After in vitro experiments showed that imipramine blue effectively inhibited movement of several cancer cell lines, the researchers tested the compound in an animal model of aggressive cancer that exhibited attributes similar to a human brain tumor called glioblastoma.
“There were many reasons why we chose to use the RT2 astrocytoma rat model for these experiments,” said Brat. “The tumor exhibited properties of aggressive growth, invasiveness, angiogenesis and necrosis that are similar to human glioblastoma; the model utilized an intact immune system, which is seen in the human disease; and the model enabled increased visualization by MRI because it was a rat model, rather than a mouse.”
Because imipramine blue is hydrophobic and doxorubicin is cytotoxic, the researchers encapsulated each compound in an artificially-prepared vesicle called a liposome so that the drugs would reach the brain. The liposomal drug delivery vehicle also ensured that the drugs would not be released into tissue until they passed through leaky blood vessel walls, which are only present where a tumor is growing.
Animals received one of the following four treatments: liposomes filled with saline, liposomes filled with imipramine blue, liposomes filled with doxorubicin chemotherapy, or liposomes filled with imipramine blue followed by liposomes filled with doxorubicin chemotherapy.
All of the animals that received the sequential treatment of imipramine blue followed by doxorubicin chemotherapy survived for 200 days -- more than 6 months -- with no observable tumor mass. Of the animals treated with doxorubicin chemotherapy alone, 33 percent were alive after 200 days with a median survival time of 44 days. Animals that received capsules filled with saline or imipramine blue – but no chemotherapy -- did not survive more than 19 days.
“Our results show that the increased effectiveness of the chemotherapy treatment is not because of a synergistic toxicity between imipramine blue and doxorubicin. Imipramine blue is not making the doxorubicin more toxic, it’s simply stopping the movement of the cancer cells and containing the cancer so that the chemotherapy can do a better job,” explained Bellamkonda, who is also the Carol Ann and David D. Flanagan Chair in Biomedical Engineering and a Georgia Cancer Coalition Distinguished Cancer Scholar.
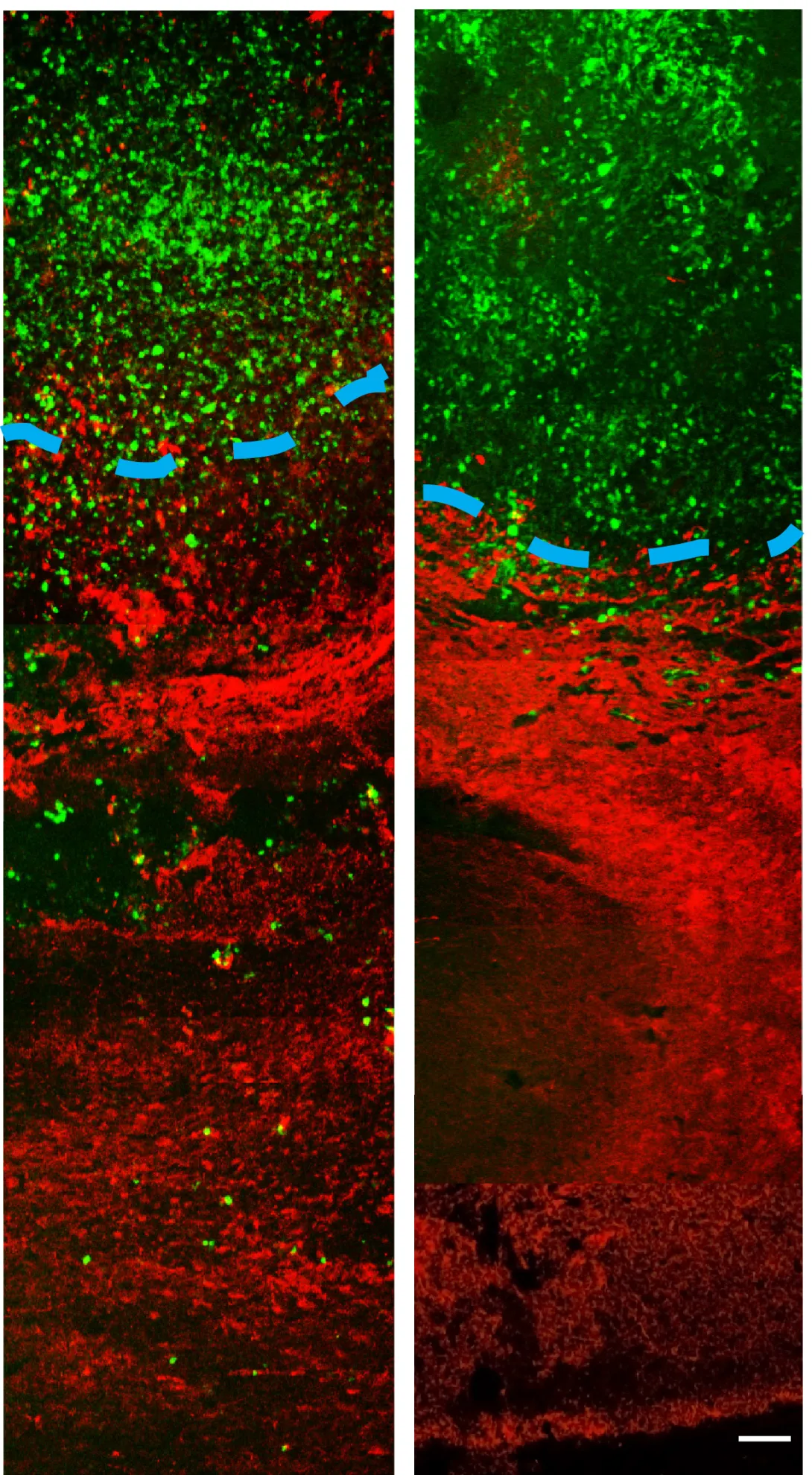
MRI results showed a reduction and compaction of the tumor in animals treated with imipramine blue followed by doxorubicin chemotherapy, while animals treated with chemotherapy alone presented with abnormal tissue and glioma cells. MRI also indicated that the blood-brain barrier breach often seen during tumor growth was present in the animals treated with chemotherapy alone, but not the group treated with chemotherapy and imipramine blue.
According to the researchers, imipramine blue appears to improve the outcome of brain cancer treatment by altering the regulation of actin, a protein found in all eukaryotic cells. Actin mediates a variety of essential biological functions, including the production of reactive oxygen species. Most cancer cells exhibit overproduction of reactive oxygen species, which are thought to stimulate cancer cells to invade healthy tissue. The dye’s reorganization of the actin cytoskeleton is thought to inhibit production of enzymes that produce reactive oxygen species.
“I formulated the imipramine blue compound as a triphenylmethane dye because I knew that another triphenylmethane dye, gentian violet, exhibited anti-cancer properties, and I decided to use imipramine -- a drug used to treat depression -- as the starting material because I knew it could get into the brain,” said Arbiser.
For future studies, the researchers are planning to test imipramine blue’s effect on animal models with invasive brain tumors, metastatic tumors, and other types of cancer such as prostate and breast.
“While we need to conduct future studies to determine if the effect of imipramine blue is the same for different types of cancer diagnosed at different stages, this initial study shows the possibility that imipramine blue may be useful as soon as any tumor is diagnosed, before anti-cancer treatment begins, to create a more treatable tumor and enhance clinical outcome,” noted Bellamkonda.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
abby@innovate.gatech.edu
404-385-3364
Feb. 13, 2012
Valentine’s Day evokes images of a stylized heart shape, but for a group of Georgia Institute of Technology researchers, the heart is a complex organ that interests them throughout the year.
Georgia Tech researchers are developing new ways to diagnose and treat heart problems -- from advanced imaging techniques and guidance for drug therapies to sophisticated surgical procedures. Georgia Tech’s emphasis on translational research accelerates the pace at which new heart-related discoveries are put to use in patient care.
Improving Heart Surgery
To advance the goal of minimally invasive cardiac surgery, researchers have developed a technology that simplifies and standardizes the technique for opening and closing the beating heart during surgery.
Apica Cardiovascular, a Georgia Tech and Emory University medical device startup, licensed the technology from the two institutions. The firm recently received a $5.5 million investment to further develop the system, which will make the transapical access and closure procedure required for delivering therapeutic devices to the heart more routine for cardiac surgeons. The goal is to expand the use of surgery techniques that are less invasive and do not require stopping the heart.
With research and development support from the Coulter Foundation Translational Research Program and the Georgia Research Alliance, the company has already completed a series of pre-clinical studies to test the functionality of the device and its biocompatibility. James Greene currently serves as the CEO of the company, which has offices in Galway, Ireland, and in Atlanta.
For more information on this work, visit http://gtresearchnews.gatech.edu/apica-cardiovascular/.
Diagnosing Heart Disease
Levent Degertekin is designing tiny devices micromachined from silicon that may make diagnosing and treating coronary artery diseases easier.
Degertekin, the George W. Woodruff Chair in Mechanical Systems, and Paul Hasler, a professor in the School of Electrical and Computer Engineering at Georgia Tech, micromachined intravascular ultrasound imaging arrays with integrated electronics. Placed on catheters inserted into the body, the devices image the arteries of the heart in three dimensions at high resolution using high-frequency ultrasound waves.
The system boasts a more compact design and three-dimensional imaging capability for guiding cardiologists during interventions, such as those for completely blocked arteries. The technology also offers higher resolution than current intravascular ultrasound systems, which help diagnose vulnerable plaque, a leading cause of heart attacks.
Funding for this research currently is provided by the National Institutes of Health. To commercialize the technology, the researchers have formed a startup company called SIBUS Medical, which is receiving assistance from VentureLab, a unit of Georgia Tech’s Enterprise Innovation Institute that nurtures faculty startup companies.
Detecting and Treating Atherosclerosis
With a five-year $14.6 million contract from the National Institutes of Health (NIH), Georgia Tech and Emory University researchers are developing nanotechnology and biomolecular engineering tools and methodologies for detecting and treating atherosclerosis. The award supports the interdisciplinary Center for Translational Cardiovascular Nanomedicine, which is led by Gang Bao, the Robert A. Milton Chair in Biomedical Engineering in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University.
Atherosclerosis typically occurs in branched or curved regions of arteries where plaques form because of cholesterol build-up. Inflammation can alter the structure of plaques so they become more likely to rupture, potentially causing a blood vessel blockage and leading to heart attack or stroke.
The researchers are working to accomplish four goals:
- Using nanoparticle probes to image and characterize atherosclerotic plaques
- Diagnosing cardiovascular disease from a blood sample
- Designing new methods for delivering anti-atherosclerosis drugs and genes into the body
- Developing stem cell based therapies to repair damaged heart tissue
Additional researchers from the Coulter Department and from Emory University are also contributing to the project. For more information on this work, visit http://gtresearchnews.gatech.edu/cardiovascular-nanomedicine-center/.
Improving Drug Dosing Following a Heart Attack
A research team led by Georgia Tech mechanical engineering assistant professor Craig Forest is designing a device to quickly and accurately personalize a patient’s drug dosage to prevent blood clots that can cause heart attacks.
When someone experiencing heart attack symptoms arrives at an emergency room, he or she typically receives a standard dose of aspirin and/or clopidogrel to prevent further blood clotting. But that standard dose may not be the best dose for a given individual.
With Forest’s device, a small blood sample is sent through a microchip containing a network of microfabricated capillaries that mimic the branching coronary arteries around the human heart. Because the branches contain flow restrictions of different sizes, the failure of blood to flow through the branches with smaller restrictions indicates that a higher drug dose may be required.
Determining the necessary dose of anti-clotting drugs can be difficult. Too much of the drug may cause the patient to experience gastrointestinal bleeding. Too little drug may allow additional clot formation and set the stage for another heart attack. Forest’s device should help determine the right dosage for each patient.
Emory University Department of Emergency Medicine assistant professor Jeremy Ackerman and Georgia Tech Regents’ professor of mechanical engineering David Ku are working with Forest on this project, which is supported by the American Heart Association.
Examining Heart Valve Leakage
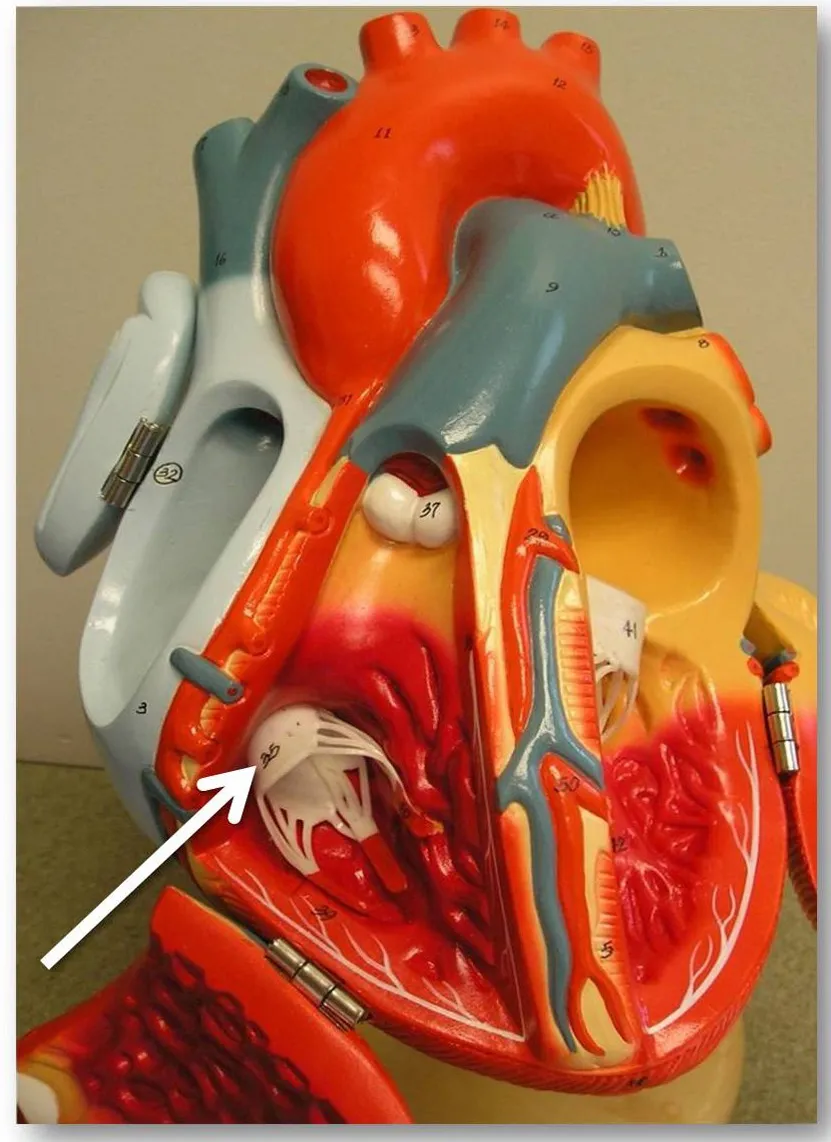
An estimated 1.6 million Americans suffer moderate to severe leakage through their tricuspid valve, a complex structure that closes off the heart’s right ventricle from the right atrium. If left untreated, severe leakage can affect an individual’s quality of life and can even lead to death.
Research teams led by Ajit Yoganathan, Georgia Tech Regents’ professor and Wallace H. Coulter Distinguished Faculty Chair in Biomedical Engineering, have discovered causes for the tricuspid valve’s leakage and ways to predict the severity of leakage in the valve. These study results could lead to improved diagnosis and treatment of the condition.
A study published in the journal Circulation found that either dilating the tricuspid valve opening or displacing the papillary muscles that control its operation can cause the valve to leak. A combination of the two actions can increase the severity of the leakage, which is called tricuspid regurgitation.
Standard clinical procedures that detail when and how tricuspid valve repairs should be performed need to be developed and this study suggests several items that should be considered in developing those protocols, according to the researchers.
In another study published in the journal Circulation: Cardiovascular Imaging, researchers found that the anatomy of the heart’s tricuspid valve can be used to predict the severity of leakage in the valve. Using 3-D echocardiograms from 64 individuals who exhibited assorted grades of tricuspid leakage, the researchers found that pulmonary arterial pressure, the size of the valve opening and papillary muscle position measurements could be used to predict the severity of an individual’s tricuspid regurgitation.
The study will change the focus and direction of future surgical therapies for tricuspid regurgitation to make them better and more durable, the researchers said.
Researchers from the Coulter Department, Emory University, Children’s Hospital Boston and Mount Sinai Medical Center contributed to these two studies.
For more information on this work, visit http://gtresearchnews.gatech.edu/tricuspid-valve-leakage/ and http://gtresearchnews.gatech.edu/tricuspid-regurgitation/.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
abby@innovate.gatech.edu
404-385-3364
Sep. 29, 2011
The National Science Foundation (NSF) has awarded $6 million to fund three projects involving researchers from the Georgia Institute of Technology. Each four-year, $2 million grant was awarded through the NSF's Division of Emerging Frontiers in Research and Innovation (EFRI).
"The EFRI research teams will probe some profound aspects of the interface of biology and engineering," said Sohi Rastegar, director of EFRI. "If they are successful, the principles and theories uncovered in their investigations could unlock many technological opportunities."
This year, 14 transformative, fundamental research projects were awarded EFRI grants in two emerging areas: technologies that build on understanding of biological signaling, and machines that can interact and cooperate with humans.
The three Georgia Tech projects include:
- Developing a "therapeutic robot" to help rehabilitate and improve motor skills in people with mobility problems;
- Creating wearable sensors that allow blind people to "see" with their hands, bodies or faces;
- Generating and rigorously testing quantitative models that describe spatial and temporal regulation of cell differentiation in tissues.
The therapeutic robot could enhance, assist and improve motor skills in humans with varying motor capabilities and deficits. The goal of the project is to program a humanoid rehabilitation robot to perform a "partnered box step," which is a defined pattern of weight shifts and directional changes, solely based on interpreting movement cues from subtle changes in forces between the hands and arms of the robot and the person.
To do this, researchers at Georgia Tech and Emory University will study how humans use their muscles to walk, balance and generate force signals with the hands for guidance when moving in cooperation with another person. They will also study "rehabilitative partnered dance," which has been specifically adapted to help improve gait and balance in individuals with motor impairments.
"Our vision is to develop robots that will interact with humans as both assistants and movement therapists," explained principal investigator Lena Ting, an associate professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. "We expect our project to have a long-term impact on quality of life of individuals with movement difficulties, such as those caused by Parkinson's disease, stroke and injury by improving fitness, motor skills and social engagement."
Working with Ting on the project are Emory University School of Medicine (geriatrics) assistant professor Madeleine Hackney, Coulter Department of Biomedical Engineering assistant professor Charlie Kemp and Georgia Tech School of Interactive Computing assistant professor Karen Liu.
For the second project, researchers at Georgia Tech and The City College of New York will investigate devices for "alternative perception" and the principles underlying the human-machine interaction. Alternative perception combines electronics and the other senses to emulate vision. In addition to aiding the visually impaired, the findings are expected to have other applications, such as the development of intelligent robots.
The researchers plan to untangle how humans learn to coordinate input from their senses -- e.g. vision, touch -- with movements, like reaching for a glass or moving through a crowded room. They will then map out how machines, such as robots and computers, learn similar tasks, to model devices that can assist humans.
The team envisions a multifunctional array of sensors on the body and has already developed prototypes for some of the devices. The full complement of wearable sensors would help a sightless person navigate by conveying information about his or her surroundings.
The researchers hope their findings on perception, and the prototypes they develop, will spawn a raft of wearable electronic devices to help blind people "see" their environment at a distance through touch, hearing and other senses. The technology would also benefit sighted individuals who must navigate in poor visibility, such as firefighters and pilots.
Principal investigator Zhigang Zhu, professor of computer science and computer engineering in City College's Grove School of Engineering, will collaborate with City College professor of psychology and director of the Program in Cognitive Neuroscience Tony Ro, City College professor of electrical engineering Ying Li Tian, Georgia Tech Woodruff School of Mechanical Engineering professor Kok-Meng Lee, and Georgia Tech School of Applied Physiology associate professor Boris Prilutsky.
The third project will address a fundamental question of developmental biology: what controls the spatial and temporal patterns of cell differentiation? Answering this question will lead to a better understanding of the basic principles of embryogenesis, explain origins of developmental disorders, and provide guidelines for tissue engineering and regenerative medicine.
The research will be conducted by principal investigator and Princeton University Department of Chemical and Biological Engineering associate professor Stanislav Shvartsman, Georgia Tech School of Chemical and Biomolecular Engineering associate professor Hang Lu, New York University Department of Biology professor Christine Rushlow, and University of Illinois at Urbana Champaign Department of Computer Science associate professor Saurabh Sinha.
Scientists know that among an embryo's first major developments is the establishment of its dorsoventral axis, which runs from its back to its belly. The researchers plan to study how this axis development unfolds -- specifically the presence and location of proteins during the process, which give rise to muscle, nerve and skin tissues.
To enable large-scale quantitative analyses of protein positional information along the dorsoventral axis, Lu and Shvartsman will further develop a microfluidic device they previously designed to reliably and robustly orient several hundred embryos in just a few minutes.
"By understanding this system at a deeper, quantitative level, we will elucidate general principles underlying the operation of genetic and multicellular networks that drive development," said Lu.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Sep. 20, 2011
The National Institutes of Health (NIH) has awarded nearly $2 million to researchers at the Georgia Institute of Technology and Emory University to develop a new class of therapeutics for treating traumatic injuries and degenerative diseases.
The five-year project focuses on developing biomaterials capable of capturing certain molecules from embryonic stem cells and delivering them to wound sites to enhance tissue regeneration in adults. By applying these unique molecules, clinicians may be able to harness the regenerative power of stem cells while avoiding concerns of tumor formation and immune system compatibility associated with most stem cell transplantation approaches.
"Pre-clinical and clinical evidence strongly suggests that the biomolecules produced by stem cells significantly impact tissue regeneration independent of differentiation into functionally competent cells," said Todd McDevitt, director of the Stem Cell Engineering Center at Georgia Tech and an associate professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. "We want to find out if the signaling molecules responsible for scarless wound healing and functional tissue restoration during early stages of embryological development can be used with adult wounds to produce successful tissue regeneration without scar formation."
In addition to McDevitt, Coulter Department associate professor Johnna Temenoff and Woodruff School of Mechanical Engineering professor Robert Guldberg are also investigators on the project.
Regenerative medicine seeks to restore normal structure and function to tissues compromised by degenerative diseases and traumatic injuries. The contrast between embryonic and adult wound healing suggests that molecules that facilitate tissue regeneration during embryonic development are distinctly different from those of adult tissues.
This grant includes plans for engineering biomaterials that can efficiently capture morphogens, which are molecules secreted by embryonic stem cells undergoing differentiation. The study will also evaluate the regenerative activity of molecule-filled biomaterials in animal models of dermal wound healing, hind limb ischemia and bone fractures. Examining the effects of the morphogens on a range of animal wound models will increase the likelihood of success and define any limitations of the technology, such as its use for specific tissues or injuries.
"Biomaterials have largely been used in an attempt to direct stem cell differentiation or serve as passive cell transplantation vehicles for regenerative medicine and tissue engineering purposes," said McDevitt, who is also a Petit Faculty Fellow in the Institute for Bioengineering and Bioscience at Georgia Tech. "The idea of specifically engineering biomaterial properties to capture and deliver complex assemblies of stem cell-derived morphogens without transplanting the cells themselves represents a novel strategy to translate the potency of stem cells into a viable regenerative medicine therapy."
The award was one of 17 granted this year through the NIH Director's Transformative Research Projects Program (T-R01), which was created to challenge the status quo with innovative ideas that have the potential to advance fields and speed the translation of research into improved health for the American public.
Another T-R01 grant was awarded to Coulter Department professor Shuming Nie, associate professor May Wang and University of Pennsylvania School of Medicine Thoracic Surgery Research Laboratory director Sunil Singhal. That $7 million, five-year grant will support continuing work by the Emory-Georgia Tech Nanotechnology Center for Personalized and Predictive Oncology team on developing fluorescent nanoparticle probes that hone in on cancer cells and on creating instruments that visualize them for cancer detection during surgery.
Since its inception in 2009, the NIH Director's Award Program has funded a total of 406 high-risk research projects, including 79 T-R01 awards.
"The NIH Director's Award programs reinvigorate the biomedical work force by providing unique opportunities to conduct research that is neither incremental nor conventional," said James M. Anderson, director of the Division of Program Coordination, Planning and Strategic Initiatives, who guides the NIH Common Fund's High-Risk Research program. "The awards are intended to catalyze giant leaps forward for any area of biomedical research, allowing investigators to go in entirely new directions."
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364
Aug. 09, 2011
A new study into the causes of leakage in one of the heart's most complex valve structures could lead to improved diagnosis and treatment of the condition.
An estimated 1.6 million Americans suffer moderate to severe leakage through their tricuspid valve, a complex structure that closes off the heart's right ventricle from the right atrium. Most people have at least some leakage in the valve, but what causes the problem is not well understood.
A new study, published online in the journal Circulation on August 1, 2011, found that either dilating the tricuspid valve opening or displacing the papillary muscles that control its operation can cause the valve to leak. A combination of the two actions can increase the severity of the leakage, which is called tricuspid regurgitation.
"We think this is the first in vitro investigation into the mechanics of the tricuspid valve, and that our findings into the mechanisms that cause tricuspid regurgitation could lead to improved diagnosis and treatment," said Ajit Yoganathan, Regents professor and Wallace H. Coulter Distinguished Faculty Chair in Biomedical Engineering in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University.
The tricuspid valve consists of three flaps that open to allow blood to flow from the heart's upper right chamber to the ventricle. To close the valve, the flaps re-cover the opening, keeping blood from flowing back into the chamber it just left. When the valve is leaky or doesn't close tightly enough, blood flows backward into the chamber just after the heart contracts.
Tricuspid regurgitation has been increasingly recognized as a source of disease in patients with chronic mitral valve regurgitation, but surgical repair of the tricuspid valve alone is recommended only in rare cases. If an individual suffers from severe tricuspid regurgitation, surgeons will sometimes repair the tricuspid valve during a surgery to repair other leaky heart valves.
"Standard clinical procedures that detail when and how tricuspid valve repairs should be performed need to be developed and this study suggests several items that should be considered," said one of the study’s co-authors David H. Adams, chair of the Department of Cardiothoracic Surgery at Mount Sinai Medical Center in New York City. "Current repairs for tricuspid regurgitation focus mainly on remodeling the valve's opening to correct enlargement, but this study shows that it may also be important to restore the position of the papillary muscles, providing as much overlap as possible, in order to conduct effective and durable tricuspid valve repairs."
With funding from the American Heart Association, Yoganathan and Coulter Department graduate student Erin Spinner conducted experiments with porcine tricuspid valves to determine possible causes of tricuspid regurgitation. The valves were attached to a plate designed to create physiological dilation and then placed inside a right heart simulator.
The researchers first investigated the effect of dilating the opening of the tricuspid valves. When the openings stretched to an area 40 percent larger than their normal size, a hole appeared in the central region of the valve. The hole caused leakage, and increased in size with further dilatation. This finding surprised the researchers because similar studies using the same method had shown that the heart's mitral valve could withstand dilation of 75 percent before leaking.
"These results tell us that the tricuspid valve is a much more complex valve than the mitral valve, which only has two flaps," explained Spinner. "Forming a proper seal over the valve opening might be more difficult with three flaps, which could be why such a large percentage of the population experiences some level of tricuspid regurgitation and why some individuals with annular dilation have tricuspid regurgitation and others do not."
The research team also investigated the effect of displacing the valve's three papillary muscles, which attach to the valve via threads. Contraction of the papillary muscles opens the valve and relaxation of the muscles closes the valve. The study showed that papillary displacement alone resulted in significant tricuspid leakage.
"While isolated displacement of the papillary muscles is rare, these results are relevant toward understanding what may happen if the size of the valve opening is repaired, but the papillary muscles are left displaced," noted Yoganathan.
The study also showed that higher levels of tricuspid leakage resulted when the researchers combined the conditions -- dilation of 40 percent or greater and displacement of all papillary muscles.
In their future work, the Coulter Department researchers plan to look at the effect of pulmonary hypertension on the tricuspid valve, because tricuspid regurgitation usually develops in association with pulmonary hypertension -- which is abnormally high blood pressure in the lungs. They also plan to work with their clinical collaborators to extend their findings to humans.
"In our in vitro study we were able to select specific porcine valves, but with human subjects there will be more anatomical variety. For example, two people may have valves of the same diameter, but one person may have longer flaps that are able to compensate for dilation whereas the other might not," noted Yoganathan.
In addition to those already mentioned, Pedro del Nido, a professor in the Department of Cardiothoracic Surgery at Children's Hospital Boston; Emir Veledar, an assistant professor with a joint appointment in the Division of Cardiology and Division of Epidemiology at Emory University; and Coulter Department research engineer Jorge Jimenez and undergraduate students Patrick Shannon and Dana Buice also contributed to this work.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364
Jul. 18, 2011
A new family of contrast agents that sneak into bacteria disguised as glucose food can detect bacterial infections in animals with high sensitivity and specificity. These agents -- called maltodextrin-based imaging probes -- can also distinguish a bacterial infection from other inflammatory conditions.
"These contrast agents fill the need for probes that can accurately image small numbers of bacteria in vivo and distinguish infections from other pathologies like cancer," said Niren Murthy, an associate professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. "These probes could ultimately improve the diagnosis and treatment of bacterial infections, which remains a major challenge in medicine."
The imaging probes were described in the July 17, 2011 advance online edition of the journal Nature Materials. The research was sponsored by the National Science Foundation and National Institutes of Health.
Coulter Department postdoctoral fellows Xinghai Ning and Seungjun Lee led the project. University of Georgia Complex Carbohydrate Research Center postdoctoral associate Zhirui Wang; and Georgia State University Department of Biology associate professor Eric Gilbert and student Bryan Subblefield also contributed to the work.
In the United States in 2010, bacterial infections caused 40,000 deaths from sepsis and were the leading cause of limb amputations. A major limitation preventing the effective treatment of bacterial infections is an inability to detect them inside the body with accuracy and sensitivity. To image bacterial infections, probes must first deliver a large quantity of the contrast agent into bacteria.
"Most existing imaging probes target the bacterial cell wall and cannot access the inside of the bacteria, but maltodextrin-based imaging probes target a bacterial ingestion pathway, which allows the contrast agent to reach a high concentration within bacteria," said Murthy.
Maltodextrin-based imaging probes consist of a fluorescent dye linked to maltohexaose, which is a major source of glucose for bacteria. The probes deliver the contrast agent into bacteria through the organism's maltodextrin transporter, which only exists in bacterial cells and not mammalian cells.
"To our knowledge, this represents the first demonstration of a targeting strategy that can deliver millimolar concentrations of an imaging probe within bacteria," noted Murthy.
In experiments using a rat model, the researchers found that the contrast agent accumulated in bacteria-infected tissues, but was efficiently cleared from uninfected tissues. They saw a 42-fold increase in fluorescence intensity between bacterial infected and uninfected tissues. However, the contrast agent did not accumulate in the healthy bacterial microflora located in the intestines. Because systemically administered glucose molecules cannot access the interior of the intestines, the bacteria located there never came into contact with the probe.
They also found that the probes could detect as few as one million viable bacteria cells. Current contrast agents for imaging bacteria require at least 100 million bacteria, according to the researchers.
In another experiment, the researchers found that the maltodextrin-based probes could distinguish between bacterial infections and inflammation with high specificity. Tissues infected with E. coli bacteria exhibited a 17-fold increase in fluorescence intensity when compared with inflamed tissues that were not infected.
Additional laboratory experiments showed that the probes could deliver large quantities of imaging probes to gram-positive and gram-negative bacteria for internalization. Both types of bacteria internalized the maltodextrin-based probes at a rate three orders of magnitude faster than mammalian cells.
"Maltodextrin-based probes show promise for imaging infections in a wide range of tissues, with an ability to detect bacteria in vivo with a sensitivity two orders of magnitude higher than previously reported," said Murthy.
This project is supported by the National Science Foundation (NSF) (NSF Career Award No. BES-0546962) and the National Institutes of Health (NIH) (Award Nos. RO1 HL096796-01 and HHSN268201000043C). The content is solely the responsibility of the principal investigator and does not necessarily represent the official views of the NSF or NIH.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Vogel Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Vogel Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364
Jun. 16, 2011
Stem cell technologies have been proposed for cell-based diagnostics and regenerative medicine therapies. However, being able to make stem cells efficiently develop into a desired cell type -- such as muscle, skin, blood vessels, bone or neurons -- limits the clinical potential of these technologies.
New research presented on June 16, 2011 at the annual meeting of the International Society for Stem Cell Research (ISSCR) shows that systematically controlling the local and global environments during stem cell development helps to effectively direct the process of differentiation. In the future, these findings could be used to develop manufacturing procedures for producing large quantities of stem cells for diagnostic and therapeutic applications. The research is sponsored by the National Science Foundation and the National Institutes of Health.
"Stem cells don't make any decisions in isolation; their decisions are spatially and temporally directed by biochemical and mechanical cues in their environment," said Todd McDevitt, director of the Stem Cell Engineering Center at Georgia Tech and an associate professor in the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University. "We have designed systems that allow us to tightly control these properties during stem cell differentiation, but also give us the flexibility to introduce a new growth factor or shake the cells a little faster to see how changes like these affect the outcome."
These systems can also be used to compare the suitability of specific stem cell types for a particular use.
"We have developed several platforms that will allow us to conduct head-to-head studies with different kinds of stem cells to determine if one type of stem cell outperforms another type for a certain application," said McDevitt, who is also a Petit Faculty Fellow in the Institute for Bioengineering and Bioscience at Georgia Tech.
Many laboratory growth methods allow stem cells to aggregate in three-dimensional clumps called "embryoid bodies" during differentiation. McDevitt and biomedical engineering graduate student Andres Bratt-Leal incorporated biomaterial particles directly within these aggregates during their formation. They introduced microparticles made of gelatin, poly(lactic-co-glycolic acid) (PLGA) or agarose and tested their impact on the assembly, intercellular communication and morphogenesis of the stem cell aggregates under different conditions by varying the microsphere-to-cell ratio and the size of the microspheres.
The researchers found that the presence of the biomaterials alone modulated embryoid body differentiation, but did not adversely affect cell viability. Compared to typical delivery methods, providing differentiation factors -- retinoic acid, bone morphogenetic protein 4 (BMP4) and vascular endothelial growth factor (VEGF) -- via microparticles induced changes in the gene and protein expression patterns of the aggregates.
By including tiny magnetic particles into the embryoid bodies during formation, the researchers also found they could use a magnet to spatially control the location of an aggregate and its assembly with other aggregates. The magnetic particles remained entrapped within the aggregates for the duration of the experiments but did not adversely affect cell viability or differentiation.
"With biomaterial and magnetic microparticles, we are beginning to be able to recreate the types of complex geometric patterns seen during early development, which require multiple cues at the same time and the ability to spatially and temporally control their local presentation," noted McDevitt.
While microparticles can be used to control differentiation by regulating the local environment, other methods exist to control differentiation through the global environment. Experiments by McDevitt and biomedical engineering graduate student Melissa Kinney have demonstrated that modulating hydrodynamic conditions can dictate the morphology of cell aggregate formation and control the expression of differentiated phenotypic cell markers.
"Because bioreactors typically impose hydrodynamic forces on cells to cultivate large volumes of cells at high density, our use of hydrodynamics to control cell fate decisions represents a novel, yet simple, principle that could be used in the future for the scalable efficient production of stem cells," added McDevitt.
Technologies capable of being directly integrated into bioprocessing systems will be the best choice for manufacturing large batches of stem cells, he noted. In the future, the development of multi-scale techniques that combine different levels of control -- both local and global -- to regulate stem cell differentiation may help the translation of stem cells into viable clinical therapies.
This project is supported by the National Science Foundation (NSF) (Award No. CBET 0651739) and the National Institutes of Health (NIH) (R01GM088291). The content is solely the responsibility of the principal investigator and does not necessarily represent the official views of the NSF or NIH.
Research News & Publications Office
Georgia Institute of Technology
75 Fifth Street, N.W., Suite 314
Atlanta, Georgia 30308 USA
Media Relations Contacts: Abby Robinson (abby@innovate.gatech.edu; 404-385-3364) or John Toon (jtoon@gatech.edu; 404-894-6986)
Writer: Abby Robinson
News Contact
Abby Robinson
Research News and Publications
Contact Abby Robinson
404-385-3364