Apr. 20, 2026
How did the earliest life on Earth build complex biological machinery with so few tools? A new study explores how the simplest building blocks of proteins — once limited to just half of today’s amino acids — could still form the sophisticated structures life depends on.
The paper, The Borderlands of Foldability: Lessons from Simplified Proteins, is a meta-analysis of six decades of protein research and reveals that ancient proteins may have been far more complicated and dynamic than previously thought.
Recently published in the journal Trends in Chemistry, the study includes Georgia Tech researchers Lynn Kamerlin, professor in the School of Chemistry and Biochemistry and Georgia Research Alliance Vasser-Woolley Chair in Molecular Design, and Quantitative Biosciences Ph.D. candidate Alfie-Louise Brownless.
Co-authors also include Institute of Science Tokyo graduate student Koh Seya and Liam M. Longo, who serves as a specially appointed associate professor at Science Tokyo and as an affiliate research scientist at the Blue Marble Space Institute of Science.
The research has implications ranging from the origins of life and the search for life in the universe to cutting-edge medical innovation. “One of the biggest unanswered questions in science is how life first began,” says Kamerlin, who is a corresponding author of the study. “Understanding how the first protein-like molecules formed and what the earliest proteins may have been like is a key part of that puzzle.”
“Proteins power our bodies — and all life on Earth,” she adds. “Simply put, the evolution of proteins is the reason that we’re able to have this conversation at all.”
A Protein Folding Paradox
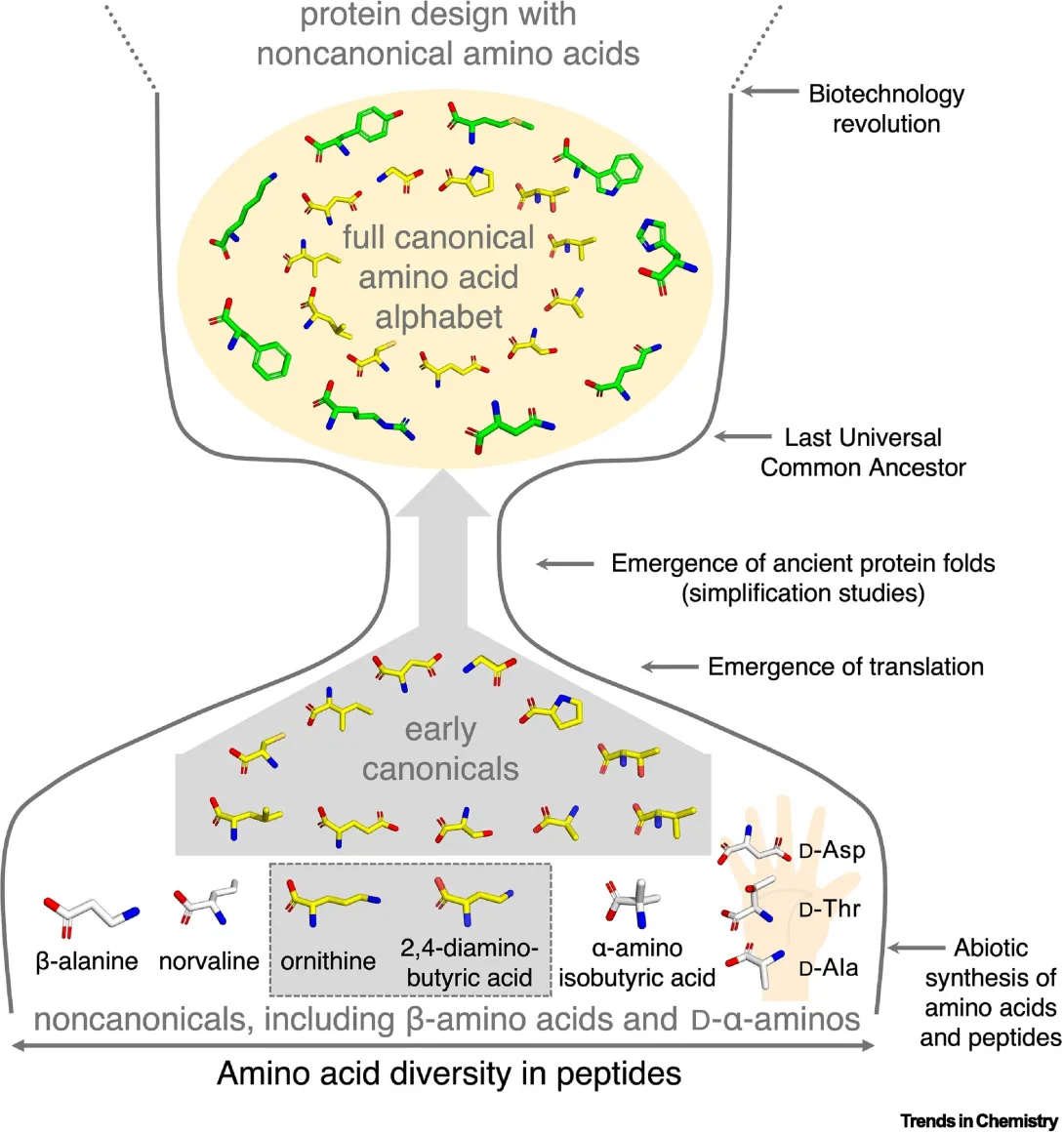
If proteins are the scaffolding of life, amino acids are the components that make up that scaffolding. “Today, an average protein is constructed from a chain of about 300 amino acids, involving 20 different types of amino acids,” Kamerlin shares. Proteins fold when these chains twist into a specific 3-dimensional shape, creating structures critical for biology.
However, while these folds are essential, exactly how a protein knows which way to fold remains a mystery. “We know that proteins didn’t just fold randomly,” Kamerlin shares, “because randomly trying all possible configurations would take a protein longer than the age of the universe.”
It’s a cornerstone problem in biological science called “Levinthal’s Paradox,” and highlights a fundamental mystery: Proteins fold incredibly quickly into very specific combinations — but like a sheet of paper spontaneously folding into an origami swan, researchers don’t know how proteins “choose” the folds they make.
“We can predict what a protein will look like, but can’t tell you how it got there,” Kamerlin adds. “That’s what we’re interested in exploring: how small early proteins developed into the complex proteins that support every living thing on today’s Earth.”
Simple Letters, Sophisticated Structures
Early proteins likely had access to just half of today’s amino acids. “About 10-12 amino acids were likely available on early Earth,” Kamerlin says. Like writing a story with just the letters “A” through “L,” researchers assumed that the ‘vocabulary’ proteins could build from such a limited amino acid alphabet would also be constrained.
“There is a language to protein folding,” Kamerlin explains. “That language is hidden in their structures. Our research is in trying to understand the rules — the grammar and vocabulary that dictate a protein fold.”
The grammar they discovered was surprising: with a combination of creative techniques and environmental support, complex structures can arise from limited amino acid alphabets.
“We found that it is possible to develop complex folds with very simple tools — and certain environments, like salty ones, can help support that,” Kamerlin shares. “Early proteins could also cross-link and associate, interacting like LEGO blocks to create more complex structures.”
Pioneering Proteins
Now, the team is conducting research in environments that could mimic conditions on early Earth — aiming to discover more about how these regions could have given rise to today’s complex proteins. “This aspect of our research also ties into the amazing space research happening at Georgia Tech,” Kamerlin says. “While we’re interested in understanding early life on Earth, our work could help inform where best to look for evidence of life beyond our planet.”
Kamerlin specializes in creating computer models that simulate possible scenarios – creating an opportunity to quickly and efficiently test many theories. The most compelling of these can then be tested by her collaborator and co-author at Science Tokyo, Liam Longo, in lab experiments.
Protein folding is also at the forefront of medical innovation, ranging from diagnostic tools to cancer treatments and neurodegenerative diseases. “In the broader scope, we’re interested in discovering what we can design, what we can stress test, and what we can reconstruct with AI and other computational tools,” Kamerlin says. “Because if you can understand how proteins fold, you gain the ability to design them.”
Funding: NASA, the Human Frontier Science Program, and the Knut and Alice Wallenberg Foundation